You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003536_00453
Basic Information
help
| Species |
|
| Lineage |
Bacteria; Verrucomicrobiota; Verrucomicrobiae; Opitutales; CAG-312; CAG-312;
|
| CAZyme ID |
MGYG000003536_00453
|
| CAZy Family |
GH28 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 379 |
|
41547.53 |
8.7069 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000003536 |
1804941 |
MAG |
Fiji |
Oceania |
|
| Gene Location |
Start: 908;
End: 2047
Strand: +
|
No EC number prediction in MGYG000003536_00453.
| Family |
Start |
End |
Evalue |
family coverage |
| GH28 |
113 |
339 |
1.2e-19 |
0.6369230769230769 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| COG5434
|
Pgu1 |
6.30e-10 |
6 |
339 |
72 |
397 |
Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam13229
|
Beta_helix |
0.010 |
148 |
276 |
2 |
119 |
Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. |
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
| AWO01000.1
|
7.61e-116 |
26 |
357 |
5 |
331 |
| APZ82864.1
|
8.23e-31 |
17 |
357 |
14 |
339 |
| BBZ75998.1
|
1.20e-06 |
148 |
300 |
159 |
287 |
This protein is predicted as SP
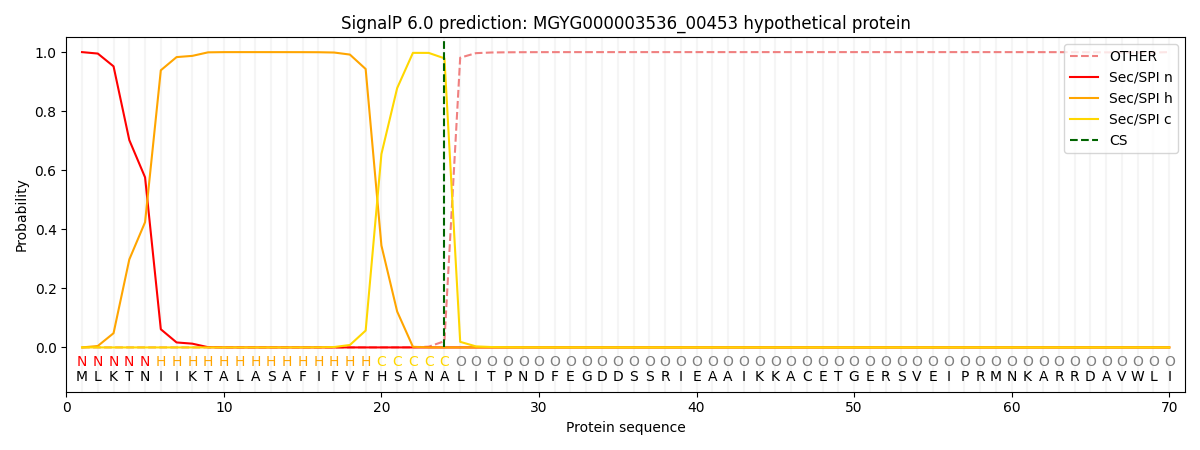
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.002440
|
0.996247
|
0.000303
|
0.000386
|
0.000305
|
0.000320
|
There is no transmembrane helices in MGYG000003536_00453.

