You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003572_00162
You are here: Home > Sequence: MGYG000003572_00162
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp900770025 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp900770025 | |||||||||||
| CAZyme ID | MGYG000003572_00162 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | Beta-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4520; End: 7444 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 22 | 741 | 2.1e-70 | 0.723404255319149 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 2.87e-20 | 23 | 440 | 9 | 414 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10340 | ebgA | 3.80e-14 | 110 | 432 | 113 | 449 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| PRK10150 | PRK10150 | 4.04e-12 | 104 | 326 | 63 | 296 | beta-D-glucuronidase; Provisional |
| PRK09525 | lacZ | 1.14e-08 | 23 | 432 | 49 | 462 | beta-galactosidase. |
| pfam02836 | Glyco_hydro_2_C | 7.56e-07 | 308 | 472 | 1 | 170 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AGB28395.1 | 0.0 | 20 | 973 | 24 | 976 |
| QVJ80649.1 | 0.0 | 7 | 973 | 2 | 900 |
| ADE82591.1 | 0.0 | 7 | 973 | 2 | 900 |
| BCS84402.1 | 0.0 | 21 | 973 | 5 | 899 |
| QUT32899.1 | 0.0 | 3 | 973 | 56 | 1014 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6XXW_A | 2.73e-19 | 102 | 434 | 68 | 418 | Structureof beta-D-Glucuronidase for Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
| 6U7J_A | 1.99e-12 | 102 | 433 | 71 | 418 | UnculturedClostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_B Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_C Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.],6U7J_D Uncultured Clostridium sp. Beta-glucuronidase [uncultured Clostridium sp.] |
| 6D8K_A | 2.64e-12 | 99 | 418 | 90 | 407 | Bacteroidesmultiple species beta-glucuronidase [Bacteroides ovatus],6D8K_B Bacteroides multiple species beta-glucuronidase [Bacteroides ovatus],6D8K_C Bacteroides multiple species beta-glucuronidase [Bacteroides ovatus],6D8K_D Bacteroides multiple species beta-glucuronidase [Bacteroides ovatus] |
| 3GM8_A | 5.07e-11 | 106 | 462 | 66 | 426 | ChainA, Glycoside hydrolase family 2, candidate beta-glycosidase [Phocaeicola vulgatus ATCC 8482] |
| 7VQM_A | 9.59e-11 | 124 | 388 | 111 | 368 | ChainA, GH2 beta-galacturonate AqGalA [Aquimarina sp.],7VQM_B Chain B, GH2 beta-galacturonate AqGalA [Aquimarina sp.],7VQM_C Chain C, GH2 beta-galacturonate AqGalA [Aquimarina sp.],7VQM_D Chain D, GH2 beta-galacturonate AqGalA [Aquimarina sp.] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KPJ7 | 1.19e-12 | 127 | 432 | 124 | 430 | Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21970 PE=2 SV=1 |
| O33815 | 8.88e-12 | 29 | 432 | 48 | 437 | Beta-galactosidase OS=Staphylococcus xylosus OX=1288 GN=lacZ PE=3 SV=1 |
| P26257 | 3.42e-10 | 95 | 426 | 43 | 382 | Beta-galactosidase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=lacZ PE=1 SV=1 |
| Q4FAT7 | 1.02e-07 | 102 | 434 | 94 | 453 | Beta-glucuronidase OS=Sus scrofa OX=9823 GN=GUSB PE=3 SV=1 |
| P05804 | 2.21e-07 | 102 | 433 | 61 | 414 | Beta-glucuronidase OS=Escherichia coli (strain K12) OX=83333 GN=uidA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
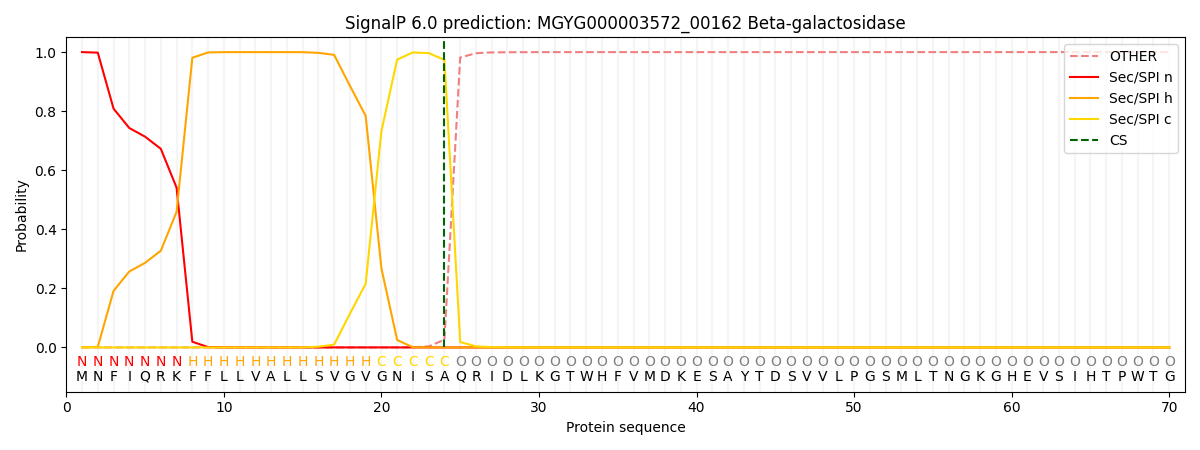
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000273 | 0.998992 | 0.000213 | 0.000165 | 0.000155 | 0.000143 |
