You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003583_01788
You are here: Home > Sequence: MGYG000003583_01788
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruminococcus_C sp900770195 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminococcus_C; Ruminococcus_C sp900770195 | |||||||||||
| CAZyme ID | MGYG000003583_01788 | |||||||||||
| CAZy Family | GH9 | |||||||||||
| CAZyme Description | Endoglucanase 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 12889; End: 15414 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH9 | 45 | 515 | 2.5e-121 | 0.992822966507177 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00759 | Glyco_hydro_9 | 4.13e-123 | 46 | 514 | 1 | 374 | Glycosyl hydrolase family 9. |
| PLN02345 | PLN02345 | 1.77e-64 | 48 | 519 | 2 | 460 | endoglucanase |
| PLN02340 | PLN02340 | 1.24e-58 | 42 | 525 | 29 | 501 | endoglucanase |
| PLN02613 | PLN02613 | 8.49e-57 | 48 | 517 | 31 | 478 | endoglucanase |
| PLN02420 | PLN02420 | 1.66e-56 | 16 | 517 | 14 | 506 | endoglucanase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL16391.1 | 1.28e-290 | 20 | 802 | 27 | 802 |
| ADU21883.1 | 1.41e-288 | 8 | 836 | 6 | 835 |
| CAP78918.2 | 2.17e-277 | 42 | 810 | 40 | 759 |
| ANV76549.1 | 8.74e-277 | 42 | 810 | 40 | 759 |
| ADU74844.1 | 8.74e-277 | 42 | 810 | 40 | 759 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2YIK_A | 5.19e-220 | 2 | 533 | 3 | 528 | ChainA, Endoglucanase [Acetivibrio thermocellus] |
| 1IA6_A | 2.58e-91 | 42 | 522 | 4 | 431 | CrystalStructure Of The Cellulase Cel9m Of C. Cellulolyticum [Ruminiclostridium cellulolyticum],1IA7_A Crystal Structure Of The Cellulase Cel9m Of C. Cellulolyticium In Complex With Cellobiose [Ruminiclostridium cellulolyticum] |
| 1G87_A | 2.19e-87 | 42 | 711 | 4 | 614 | TheCrystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum [Ruminiclostridium cellulolyticum],1G87_B The Crystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum [Ruminiclostridium cellulolyticum],1GA2_A The Crystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum Complexed With Cellobiose [Ruminiclostridium cellulolyticum],1GA2_B The Crystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum Complexed With Cellobiose [Ruminiclostridium cellulolyticum],1KFG_A The X-ray Crystal Structure of Cel9G from Clostridium cellulolyticum complexed with a Thio-Oligosaccharide Inhibitor [Ruminiclostridium cellulolyticum],1KFG_B The X-ray Crystal Structure of Cel9G from Clostridium cellulolyticum complexed with a Thio-Oligosaccharide Inhibitor [Ruminiclostridium cellulolyticum] |
| 1K72_A | 5.99e-86 | 42 | 711 | 4 | 614 | TheX-ray Crystal Structure Of Cel9G Complexed With cellotriose [Ruminiclostridium cellulolyticum],1K72_B The X-ray Crystal Structure Of Cel9G Complexed With cellotriose [Ruminiclostridium cellulolyticum] |
| 2XFG_A | 2.56e-83 | 42 | 522 | 24 | 464 | ChainA, ENDOGLUCANASE 1 [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02934 | 1.20e-93 | 42 | 720 | 76 | 692 | Endoglucanase 1 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celI PE=1 SV=2 |
| Q5YLG1 | 1.22e-91 | 41 | 712 | 46 | 658 | Endoglucanase A OS=Bacillus pumilus OX=1408 GN=eglA PE=1 SV=1 |
| P37700 | 1.55e-90 | 42 | 838 | 39 | 723 | Endoglucanase G OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCG PE=1 SV=2 |
| P28622 | 5.63e-83 | 40 | 711 | 26 | 634 | Endoglucanase 4 OS=Bacillus sp. (strain KSM-522) OX=120046 PE=3 SV=2 |
| P26224 | 1.02e-80 | 41 | 806 | 29 | 699 | Endoglucanase F OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
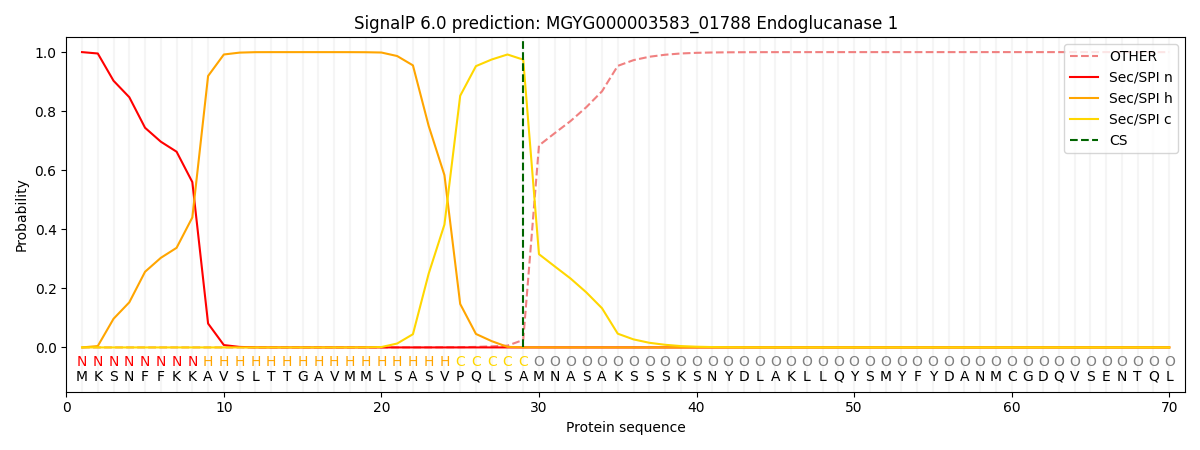
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000366 | 0.998812 | 0.000230 | 0.000215 | 0.000180 | 0.000153 |
