You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003625_01470
You are here: Home > Sequence: MGYG000003625_01470
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
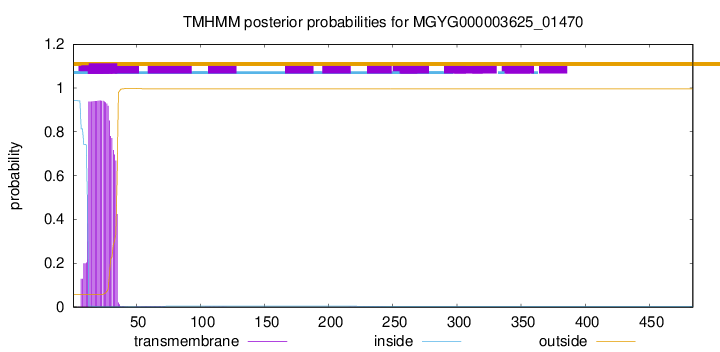
TMHMM annotations
Basic Information help
| Species | UBA2913 sp900770895 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Selenomonadales; Selenomonadaceae; UBA2913; UBA2913 sp900770895 | |||||||||||
| CAZyme ID | MGYG000003625_01470 | |||||||||||
| CAZy Family | GH84 | |||||||||||
| CAZyme Description | O-GlcNAcase NagJ | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7401; End: 8855 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH84 | 38 | 334 | 2.7e-88 | 0.9796610169491525 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07555 | NAGidase | 2.59e-113 | 38 | 329 | 1 | 284 | beta-N-acetylglucosaminidase. This family has previously been described as a hyaluronidase. However, more recently it has been shown that this family has beta-N-acetylglucosaminidase activity. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAL82559.1 | 2.23e-136 | 25 | 479 | 24 | 456 |
| AEC00383.1 | 1.58e-105 | 36 | 439 | 41 | 434 |
| ASY51512.1 | 5.21e-89 | 35 | 478 | 179 | 616 |
| AXH52449.1 | 5.21e-89 | 35 | 478 | 179 | 616 |
| BAB80940.1 | 5.21e-89 | 35 | 478 | 179 | 616 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2CBI_A | 6.17e-93 | 35 | 478 | 149 | 586 | Structureof the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase [Clostridium perfringens],2CBI_B Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase [Clostridium perfringens],2CBJ_A Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc [Clostridium perfringens],2CBJ_B Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc [Clostridium perfringens],2V5C_A Family 84 glycoside hydrolase from Clostridium perfringens, 2.1 Angstrom structure [Clostridium perfringens],2V5C_B Family 84 glycoside hydrolase from Clostridium perfringens, 2.1 Angstrom structure [Clostridium perfringens],2VUR_A Chemical dissection of the link between Streptozotocin, O-GlcNAc and pancreatic cell death [Clostridium perfringens],2VUR_B Chemical dissection of the link between Streptozotocin, O-GlcNAc and pancreatic cell death [Clostridium perfringens],2X0Y_A Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds [Clostridium perfringens],2X0Y_B Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds [Clostridium perfringens] |
| 5OXD_A | 7.84e-93 | 35 | 478 | 151 | 588 | Complexof a C. perfringens O-GlcNAcase with a fragment hit [Clostridium perfringens] |
| 2J62_A | 8.68e-93 | 35 | 478 | 149 | 586 | Structureof a bacterial O-glcnacase in complex with glcnacstatin [Clostridium perfringens],2J62_B Structure of a bacterial O-glcnacase in complex with glcnacstatin [Clostridium perfringens],2WB5_A GlcNAcstatins are nanomolar inhibitors of human O-GlcNAcase inducing cellular hyper-O-GlcNAcylation [Clostridium perfringens],2WB5_B GlcNAcstatins are nanomolar inhibitors of human O-GlcNAcase inducing cellular hyper-O-GlcNAcylation [Clostridium perfringens] |
| 4ZXL_A | 3.25e-92 | 35 | 478 | 141 | 578 | CpOGAD298N in complex with Drosophila HCF -derived Thr-O-GlcNAc peptide [Clostridium perfringens ATCC 13124] |
| 2YDQ_A | 4.31e-92 | 35 | 478 | 151 | 588 | CpOGAD298N in complex with hOGA-derived O-GlcNAc peptide [Clostridium perfringens],2YDR_A CpOGA D298N in complex with p53-derived O-GlcNAc peptide [Clostridium perfringens],2YDS_A CpOGA D298N in complex with TAB1-derived O-GlcNAc peptide [Clostridium perfringens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8XL08 | 1.04e-89 | 35 | 478 | 179 | 616 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=nagJ PE=1 SV=1 |
| Q0TR53 | 1.04e-88 | 35 | 478 | 179 | 616 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain ATCC 13124 / DSM 756 / JCM 1290 / NCIMB 6125 / NCTC 8237 / Type A) OX=195103 GN=nagJ PE=1 SV=1 |
| Q89ZI2 | 2.17e-55 | 38 | 368 | 151 | 473 | O-GlcNAcase BT_4395 OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_4395 PE=1 SV=1 |
| Q2CEE3 | 6.29e-36 | 37 | 338 | 2 | 303 | Protein O-GlcNAcase OS=Oceanicola granulosus (strain ATCC BAA-861 / DSM 15982 / KCTC 12143 / HTCC2516) OX=314256 GN=OG2516_04129 PE=1 SV=1 |
| O60502 | 1.38e-31 | 24 | 302 | 46 | 327 | Protein O-GlcNAcase OS=Homo sapiens OX=9606 GN=OGA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
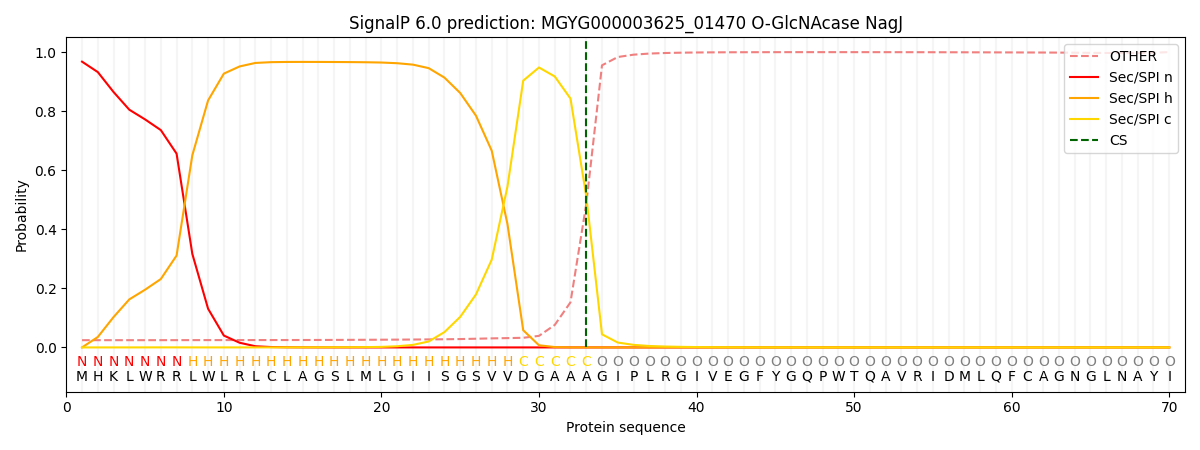
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.031917 | 0.956384 | 0.002678 | 0.008030 | 0.000538 | 0.000406 |