You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003625_01759
You are here: Home > Sequence: MGYG000003625_01759
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA2913 sp900770895 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Selenomonadales; Selenomonadaceae; UBA2913; UBA2913 sp900770895 | |||||||||||
| CAZyme ID | MGYG000003625_01759 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | Exo-poly-alpha-D-galacturonosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21173; End: 22840 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 172 | 549 | 1.7e-58 | 0.963076923076923 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 6.41e-65 | 38 | 514 | 4 | 468 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 1.12e-10 | 309 | 531 | 88 | 304 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| pfam12708 | Pectate_lyase_3 | 1.99e-10 | 150 | 258 | 3 | 128 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| PLN02218 | PLN02218 | 3.40e-10 | 146 | 513 | 65 | 391 | polygalacturonase ADPG |
| cd00063 | FN3 | 2.87e-07 | 29 | 135 | 1 | 86 | Fibronectin type 3 domain; One of three types of internal repeats found in the plasma protein fibronectin. Its tenth fibronectin type III repeat contains an RGD cell recognition sequence in a flexible loop between 2 strands. Approximately 2% of all animal proteins contain the FN3 repeat; including extracellular and intracellular proteins, membrane spanning cytokine receptors, growth hormone receptors, tyrosine phosphatase receptors, and adhesion molecules. FN3-like domains are also found in bacterial glycosyl hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AIF50363.1 | 1.01e-149 | 31 | 555 | 38 | 580 |
| BAL81848.1 | 2.77e-146 | 29 | 549 | 27 | 560 |
| AJQ29053.1 | 2.39e-137 | 16 | 552 | 18 | 579 |
| AGF59029.1 | 7.32e-129 | 31 | 552 | 38 | 562 |
| QOH53990.1 | 2.70e-125 | 3 | 551 | 8 | 575 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2UVE_A | 3.69e-108 | 31 | 551 | 37 | 582 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 3JUR_A | 2.91e-23 | 153 | 523 | 32 | 403 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 5OLP_A | 3.42e-16 | 150 | 551 | 46 | 448 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 1BHE_A | 3.91e-13 | 157 | 531 | 19 | 362 | ChainA, POLYGALACTURONASE [Pectobacterium carotovorum] |
| 4MXN_A | 2.61e-09 | 150 | 267 | 23 | 140 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P15922 | 1.34e-123 | 9 | 551 | 13 | 575 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
| P18192 | 3.34e-13 | 157 | 531 | 45 | 388 | Endo-polygalacturonase OS=Pectobacterium carotovorum subsp. carotovorum OX=555 GN=peh PE=3 SV=1 |
| P26509 | 2.48e-12 | 157 | 531 | 45 | 388 | Endo-polygalacturonase OS=Pectobacterium parmentieri OX=1905730 GN=pehA PE=1 SV=1 |
| P20041 | 1.17e-11 | 164 | 495 | 72 | 421 | Polygalacturonase OS=Ralstonia solanacearum OX=305 GN=pglA PE=1 SV=1 |
| P58598 | 2.56e-10 | 164 | 507 | 74 | 427 | Polygalacturonase OS=Ralstonia solanacearum (strain GMI1000) OX=267608 GN=pglA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
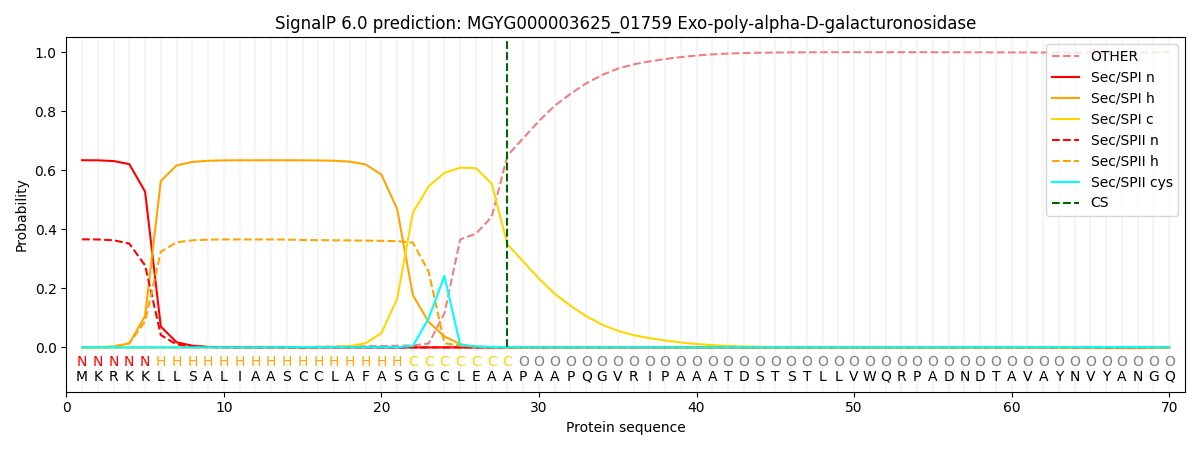
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000722 | 0.624006 | 0.374285 | 0.000464 | 0.000271 | 0.000221 |
