You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003634_00776
You are here: Home > Sequence: MGYG000003634_00776
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | SFVR01 sp900771035 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Paludibacteraceae; SFVR01; SFVR01 sp900771035 | |||||||||||
| CAZyme ID | MGYG000003634_00776 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4505; End: 5623 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 47 | 370 | 1e-34 | 0.9076923076923077 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.53e-29 | 11 | 326 | 70 | 407 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| PLN02218 | PLN02218 | 6.51e-08 | 23 | 230 | 67 | 268 | polygalacturonase ADPG |
| pfam12708 | Pectate_lyase_3 | 9.88e-07 | 24 | 76 | 2 | 59 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| PLN02793 | PLN02793 | 3.54e-06 | 22 | 345 | 51 | 365 | Probable polygalacturonase |
| pfam00295 | Glyco_hydro_28 | 1.00e-05 | 128 | 242 | 50 | 176 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIU97396.1 | 1.04e-117 | 11 | 370 | 18 | 379 |
| QDH57554.1 | 9.48e-116 | 12 | 370 | 19 | 379 |
| QJD94620.1 | 2.85e-100 | 25 | 370 | 36 | 383 |
| CRY97779.1 | 8.09e-98 | 83 | 372 | 6 | 289 |
| QUT51565.1 | 2.17e-94 | 23 | 372 | 35 | 372 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3JUR_A | 2.72e-14 | 22 | 372 | 26 | 407 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 2UVE_A | 3.61e-12 | 24 | 229 | 157 | 405 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 4MXN_A | 6.31e-12 | 17 | 265 | 15 | 235 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
| 5OLP_A | 5.05e-10 | 22 | 251 | 43 | 293 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7PZL3 | 7.13e-12 | 16 | 230 | 55 | 275 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| P35336 | 9.06e-12 | 22 | 322 | 88 | 369 | Polygalacturonase OS=Actinidia deliciosa OX=3627 PE=2 SV=1 |
| P15922 | 4.58e-10 | 22 | 229 | 150 | 398 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
| P48978 | 8.81e-09 | 22 | 365 | 97 | 420 | Polygalacturonase OS=Malus domestica OX=3750 PE=2 SV=1 |
| P49062 | 6.05e-08 | 20 | 349 | 48 | 357 | Exopolygalacturonase clone GBGE184 OS=Arabidopsis thaliana OX=3702 GN=PGA3 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
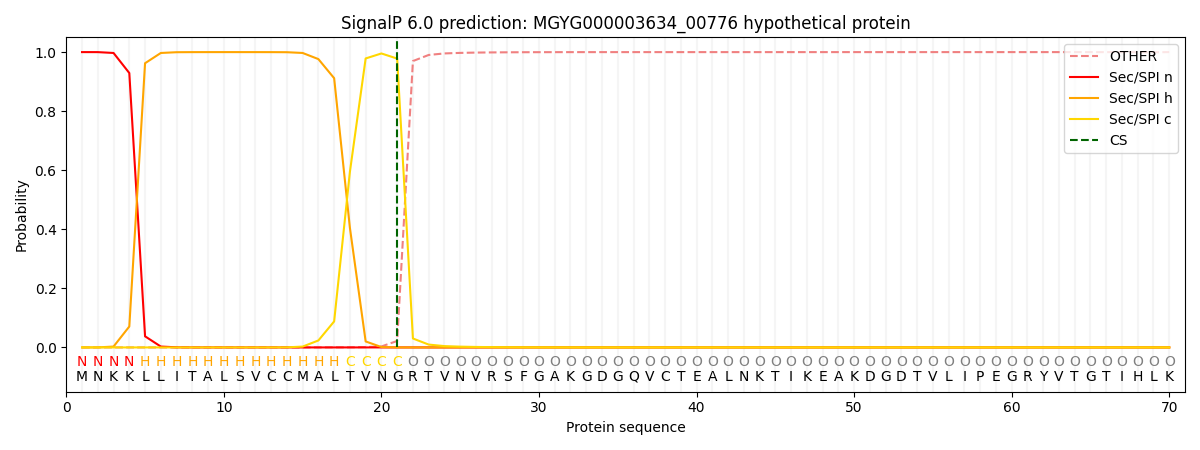
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000361 | 0.998654 | 0.000399 | 0.000187 | 0.000184 | 0.000176 |
