You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003648_02405
You are here: Home > Sequence: MGYG000003648_02405
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
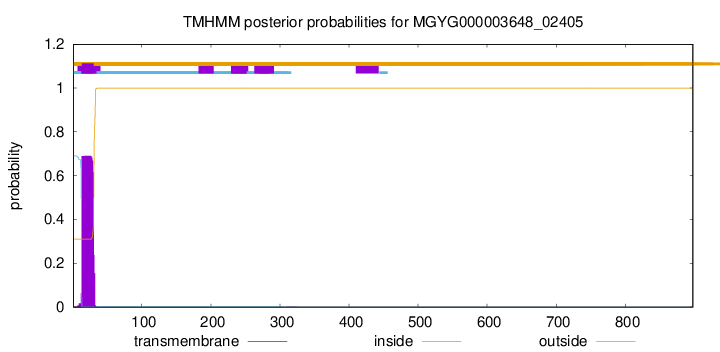
TMHMM annotations
Basic Information help
| Species | NK4A136 sp002314855 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; NK4A136; NK4A136 sp002314855 | |||||||||||
| CAZyme ID | MGYG000003648_02405 | |||||||||||
| CAZy Family | GH9 | |||||||||||
| CAZyme Description | Endoglucanase 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 33285; End: 35981 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH9 | 104 | 529 | 2.8e-133 | 0.9976076555023924 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00759 | Glyco_hydro_9 | 7.93e-154 | 107 | 528 | 1 | 374 | Glycosyl hydrolase family 9. |
| PLN02340 | PLN02340 | 5.31e-109 | 103 | 533 | 29 | 495 | endoglucanase |
| PLN00119 | PLN00119 | 2.22e-108 | 104 | 531 | 31 | 488 | endoglucanase |
| PLN02345 | PLN02345 | 9.79e-105 | 108 | 533 | 1 | 460 | endoglucanase |
| PLN02171 | PLN02171 | 2.84e-103 | 99 | 554 | 25 | 518 | endoglucanase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABN51281.1 | 5.88e-273 | 99 | 898 | 72 | 887 |
| CAB76932.1 | 5.88e-273 | 99 | 898 | 72 | 887 |
| ALX09207.1 | 1.18e-272 | 99 | 898 | 72 | 887 |
| ANV76959.1 | 1.18e-272 | 99 | 898 | 72 | 887 |
| ADU75232.1 | 1.18e-272 | 99 | 898 | 72 | 887 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1G87_A | 5.26e-251 | 101 | 704 | 2 | 614 | TheCrystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum [Ruminiclostridium cellulolyticum],1G87_B The Crystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum [Ruminiclostridium cellulolyticum],1GA2_A The Crystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum Complexed With Cellobiose [Ruminiclostridium cellulolyticum],1GA2_B The Crystal Structure Of Endoglucanase 9g From Clostridium Cellulolyticum Complexed With Cellobiose [Ruminiclostridium cellulolyticum],1KFG_A The X-ray Crystal Structure of Cel9G from Clostridium cellulolyticum complexed with a Thio-Oligosaccharide Inhibitor [Ruminiclostridium cellulolyticum],1KFG_B The X-ray Crystal Structure of Cel9G from Clostridium cellulolyticum complexed with a Thio-Oligosaccharide Inhibitor [Ruminiclostridium cellulolyticum] |
| 1K72_A | 1.71e-249 | 101 | 704 | 2 | 614 | TheX-ray Crystal Structure Of Cel9G Complexed With cellotriose [Ruminiclostridium cellulolyticum],1K72_B The X-ray Crystal Structure Of Cel9G Complexed With cellotriose [Ruminiclostridium cellulolyticum] |
| 2XFG_A | 6.22e-212 | 99 | 535 | 20 | 463 | ChainA, ENDOGLUCANASE 1 [Acetivibrio thermocellus] |
| 4DOD_A | 5.75e-207 | 99 | 547 | 22 | 475 | Thestructure of Cbescii CelA GH9 module [Caldicellulosiruptor bescii],4DOE_A The liganded structure of Cbescii CelA GH9 module [Caldicellulosiruptor bescii] |
| 1JS4_A | 1.17e-183 | 103 | 704 | 4 | 605 | EndoEXOCELLULASE:CELLOBIOSEFROM THERMOMONOSPORA [Thermobifida fusca],1JS4_B EndoEXOCELLULASE:CELLOBIOSE FROM THERMOMONOSPORA [Thermobifida fusca],1TF4_A EndoEXOCELLULASE FROM THERMOMONOSPORA [Thermobifida fusca],1TF4_B EndoEXOCELLULASE FROM THERMOMONOSPORA [Thermobifida fusca],3TF4_A EndoEXOCELLULASE:CELLOTRIOSE FROM THERMOMONOSPORA [Thermobifida fusca],3TF4_B EndoEXOCELLULASE:CELLOTRIOSE FROM THERMOMONOSPORA [Thermobifida fusca],4TF4_A EndoEXOCELLULASE:CELLOPENTAOSE FROM THERMOMONOSPORA [Thermobifida fusca],4TF4_B EndoEXOCELLULASE:CELLOPENTAOSE FROM THERMOMONOSPORA [Thermobifida fusca] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02934 | 1.18e-273 | 99 | 898 | 72 | 887 | Endoglucanase 1 OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celI PE=1 SV=2 |
| P37700 | 1.53e-250 | 99 | 708 | 35 | 653 | Endoglucanase G OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCG PE=1 SV=2 |
| P23659 | 2.14e-247 | 103 | 715 | 30 | 652 | Endoglucanase Z OS=Thermoclostridium stercorarium OX=1510 GN=celZ PE=1 SV=1 |
| P22534 | 1.83e-246 | 101 | 704 | 24 | 636 | Endoglucanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celA PE=3 SV=2 |
| P26224 | 2.40e-234 | 97 | 708 | 24 | 642 | Endoglucanase F OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
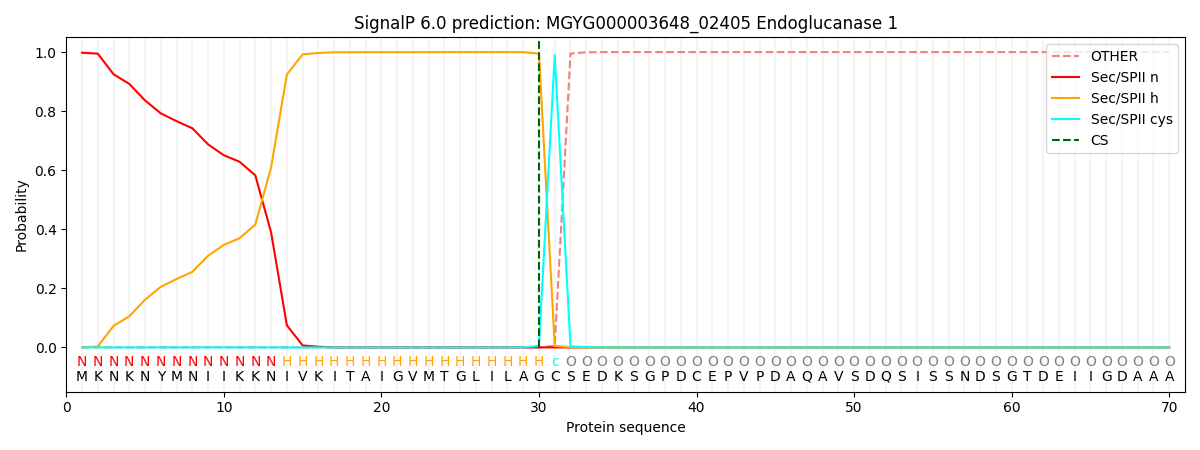
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000052 | 0.000391 | 0.999575 | 0.000007 | 0.000002 | 0.000001 |