You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003681_00880
You are here: Home > Sequence: MGYG000003681_00880
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides stercoris | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides stercoris | |||||||||||
| CAZyme ID | MGYG000003681_00880 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 168184; End: 169458 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 185 | 384 | 1.2e-33 | 0.5815384615384616 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 2.74e-14 | 111 | 392 | 144 | 406 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam00295 | Glyco_hydro_28 | 8.63e-06 | 191 | 285 | 88 | 164 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| pfam13229 | Beta_helix | 2.46e-04 | 191 | 338 | 3 | 137 | Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. |
| PLN03010 | PLN03010 | 2.77e-04 | 171 | 379 | 115 | 311 | polygalacturonase |
| pfam05048 | NosD | 5.93e-04 | 168 | 305 | 2 | 139 | Periplasmic copper-binding protein (NosD). NosD is a periplasmic protein which is thought to insert copper into the exported reductase apoenzyme (NosZ). This region forms a parallel beta helix domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ62050.1 | 1.54e-250 | 17 | 405 | 14 | 402 |
| QUT92200.1 | 3.10e-250 | 17 | 405 | 14 | 402 |
| QUU00181.1 | 8.11e-249 | 19 | 405 | 19 | 405 |
| QUT34117.1 | 1.56e-246 | 19 | 405 | 19 | 405 |
| QQY39815.1 | 5.97e-241 | 20 | 405 | 17 | 402 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A1DBR6 | 6.02e-06 | 200 | 380 | 240 | 393 | Probable endopolygalacturonase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=pgaD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
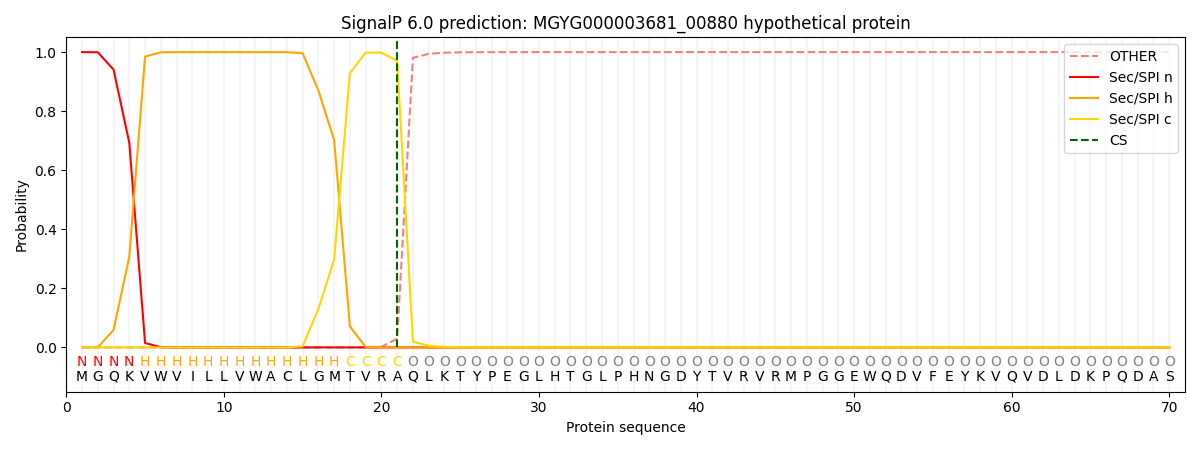
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000207 | 0.999189 | 0.000163 | 0.000138 | 0.000140 | 0.000133 |
