You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003682_02615
You are here: Home > Sequence: MGYG000003682_02615
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruthenibacterium lactatiformans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruthenibacterium; Ruthenibacterium lactatiformans | |||||||||||
| CAZyme ID | MGYG000003682_02615 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 13924; End: 15063 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 45 | 350 | 1.5e-32 | 0.9210526315789473 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 7.11e-50 | 45 | 354 | 16 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| pfam13088 | BNR_2 | 1.38e-24 | 54 | 336 | 1 | 280 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
| pfam04616 | Glyco_hydro_43 | 7.25e-04 | 37 | 119 | 63 | 133 | Glycosyl hydrolases family 43. The glycosyl hydrolase family 43 contains members that are arabinanases. Arabinanases hydrolyze the alpha-1,5-linked L-arabinofuranoside backbone of plant cell wall arabinans. The structure of arabinanase Arb43A from Cellvibrio japonicus reveals a five-bladed beta-propeller fold. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
| COG3507 | XynB2 | 0.001 | 60 | 127 | 111 | 168 | Beta-xylosidase [Carbohydrate transport and metabolism]. |
| cd18617 | GH43_XynB-like | 0.004 | 43 | 131 | 66 | 148 | Glycosyl hydrolase family 43, such as Bacteroides ovatus alpha-L-arabinofuranosidase (BoGH43, XynB). This glycosyl hydrolase family 43 (GH43) subgroup includes enzymes that have been characterized to have alpha-L-arabinofuranosidase (EC 3.2.1.55) and beta-1,4-xylosidase (beta-D-xylosidase;xylan 1,4-beta-xylosidase; EC 3.2.1.37) activities. Beta-1,4-xylosidases are part of an array of hemicellulases that are involved in the final breakdown of plant cell-wall whereby they degrade xylan. They hydrolyze beta-1,4 glycosidic bonds between two xylose units in short xylooligosaccharides. These are inverting enzymes (i.e. they invert the stereochemistry of the anomeric carbon atom of the substrate) that have an aspartate as the catalytic general base, a glutamate as the catalytic general acid and another aspartate that is responsible for pKa modulation and orienting the catalytic acid. Also included in this subfamily are Bacteroides ovatus alpha-L-arabinofuranosidases, BoGH43A and BoGH43B, both having a two-domain architecture, consisting of an N-terminal 5-bladed beta-propeller domain harboring the catalytic active site, and a C-terminal beta-sandwich domain. However, despite significant functional overlap between these two enzymes, BoGH43A and BoGH43B share just 41% sequence identity. The latter appears to be significantly less active on the same substrates, suggesting that these paralogs may play subtly different roles during the degradation of xyloglucans from different sources, or may function most optimally at different stages in the catabolism of xyloglucan oligosaccharides (XyGOs), for example before or after hydrolysis of certain side-chain moieties. It also includes Phanerochaete chrysosporium BKM-F-1767 Xyl, a bifunctional xylosidase/arabinofuranosidase. A common structural feature of GH43 enzymes is a 5-bladed beta-propeller domain that contains the catalytic acid and catalytic base. A long V-shaped groove, partially enclosed at one end, forms a single extended substrate-binding surface across the face of the propeller. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDT91705.1 | 8.16e-124 | 16 | 377 | 29 | 388 |
| QDU15387.1 | 1.64e-123 | 11 | 377 | 24 | 388 |
| QEG17440.1 | 6.60e-123 | 16 | 377 | 25 | 388 |
| QDT79764.1 | 2.66e-122 | 16 | 377 | 25 | 388 |
| QGQ32653.1 | 1.08e-121 | 42 | 377 | 2 | 336 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
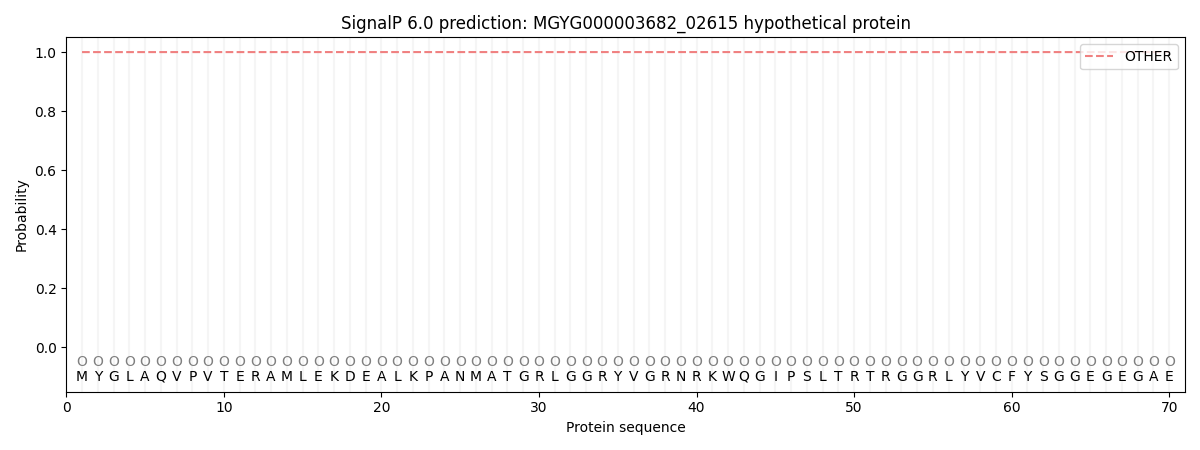
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000048 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
