You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003687_03064
You are here: Home > Sequence: MGYG000003687_03064
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
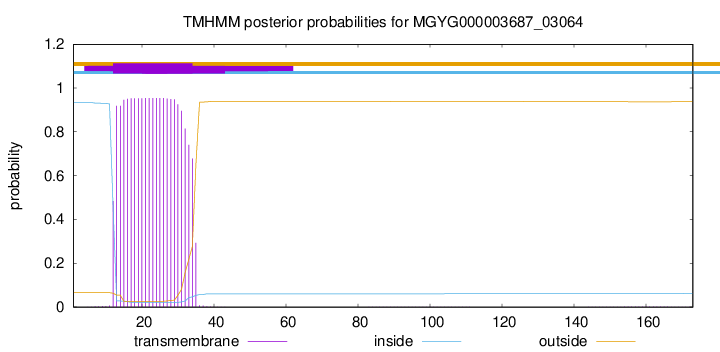
TMHMM annotations
Basic Information help
| Species | Paenibacillus polymyxa | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus; Paenibacillus polymyxa | |||||||||||
| CAZyme ID | MGYG000003687_03064 | |||||||||||
| CAZy Family | PL3 | |||||||||||
| CAZyme Description | Pectate lyase A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 110179; End: 110700 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL3 | 36 | 154 | 1.6e-53 | 0.6763005780346821 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam03211 | Pectate_lyase | 1.74e-60 | 31 | 153 | 6 | 127 | Pectate lyase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDA26106.1 | 1.26e-105 | 1 | 153 | 1 | 153 |
| AUS25157.1 | 1.26e-105 | 1 | 153 | 1 | 153 |
| QPK51959.1 | 1.26e-105 | 1 | 153 | 1 | 153 |
| AHM64591.1 | 1.26e-105 | 1 | 153 | 1 | 153 |
| AIY10231.1 | 1.26e-105 | 1 | 153 | 1 | 153 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1EE6_A | 6.97e-59 | 34 | 153 | 1 | 120 | CrystalStructure Of Pectate Lyase From Bacillus Sp. Strain Ksm-P15. [Bacillus sp. KSM-P15] |
| 4EW9_A | 7.24e-43 | 37 | 153 | 9 | 123 | Theliganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725],4EW9_B The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725] |
| 3T9G_A | 7.46e-43 | 37 | 153 | 10 | 124 | Thecrystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii],3T9G_B The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii] |
| 4Z03_A | 7.53e-42 | 37 | 153 | 18 | 132 | C.bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725],4Z03_B C. bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725] |
| 4Z06_A | 7.53e-42 | 37 | 153 | 18 | 132 | C.bescii Family 3 pectate lyase double mutant K108A/R133A in complex with ALPHA-D-GALACTOPYRANURONIC ACID [Caldicellulosiruptor bescii DSM 6725],4Z06_B C. bescii Family 3 pectate lyase double mutant K108A/R133A in complex with ALPHA-D-GALACTOPYRANURONIC ACID [Caldicellulosiruptor bescii DSM 6725] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D3JTC1 | 8.25e-63 | 9 | 153 | 1 | 145 | Pectate lyase A OS=Paenibacillus amylolyticus OX=1451 GN=pelA PE=1 SV=1 |
| Q9X6Z2 | 3.33e-62 | 9 | 153 | 1 | 145 | Pectate lyase A OS=Paenibacillus barcinonensis OX=198119 GN=pelA PE=1 SV=1 |
| Q65EF5 | 1.82e-51 | 14 | 153 | 7 | 146 | Pectate lyase C OS=Bacillus licheniformis (strain ATCC 14580 / DSM 13 / JCM 2505 / CCUG 7422 / NBRC 12200 / NCIMB 9375 / NCTC 10341 / NRRL NRS-1264 / Gibson 46) OX=279010 GN=pelC PE=3 SV=2 |
| O34310 | 1.82e-43 | 33 | 153 | 27 | 146 | Pectate lyase C OS=Bacillus subtilis (strain 168) OX=224308 GN=pelC PE=1 SV=1 |
| Q0CJ49 | 8.99e-19 | 43 | 152 | 48 | 156 | Probable pectate lyase D OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
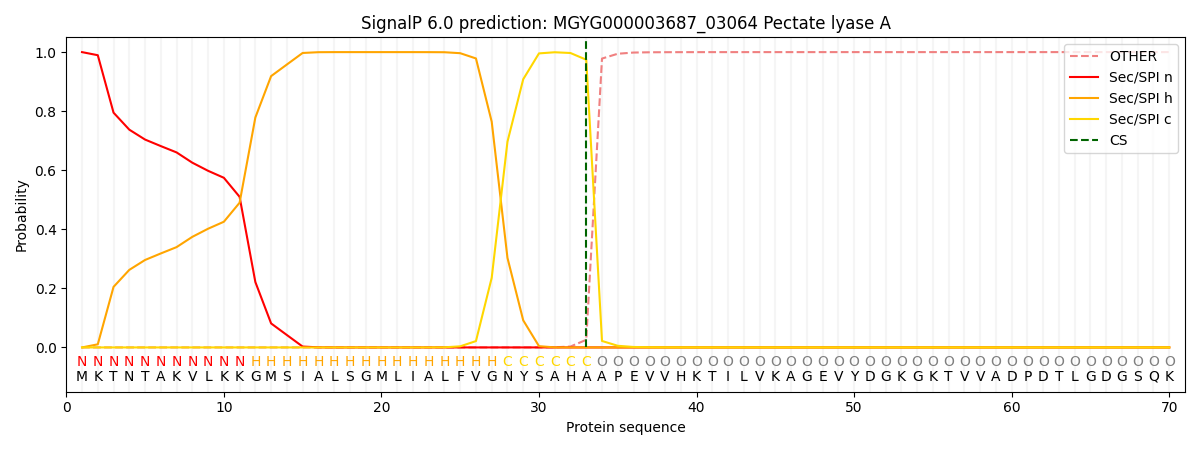
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000293 | 0.998970 | 0.000190 | 0.000187 | 0.000171 | 0.000151 |