You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003727_01137
You are here: Home > Sequence: MGYG000003727_01137
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Peptoniphilus_A lacrimalis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Tissierellales; Peptoniphilaceae; Peptoniphilus_A; Peptoniphilus_A lacrimalis | |||||||||||
| CAZyme ID | MGYG000003727_01137 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 26142; End: 29789 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4193 | LytD | 2.40e-29 | 406 | 542 | 79 | 228 | Beta- N-acetylglucosaminidase [Carbohydrate transport and metabolism]. |
| smart00047 | LYZ2 | 7.17e-23 | 418 | 541 | 1 | 138 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| pfam05382 | Amidase_5 | 2.33e-20 | 246 | 375 | 2 | 136 | Bacteriophage peptidoglycan hydrolase. At least one of the members of this family, the Pal protein from the pneumococcal bacteriophage Dp-1 has been shown to be a N-acetylmuramoyl-L-alanine amidase. According to the known modular structure of this and other peptidoglycan hydrolases from the pneumococcal system, the active site should reside at the N-terminal domain whereas the C-terminal domain binds to the choline residues of the cell wall teichoic acids. This family appears to be related to pfam00877. |
| pfam01832 | Glucosaminidase | 2.19e-14 | 430 | 475 | 1 | 46 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan. |
| TIGR01665 | put_anti_recept | 3.81e-09 | 27 | 219 | 1 | 193 | phage minor structural protein, N-terminal region. This model represents the conserved N-terminal region, typically from about residue 25 to about residue 350, of a family of uncharacterized phage proteins 500 to 1700 residues in length. [Mobile and extrachromosomal element functions, Prophage functions] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QKH79738.1 | 0.0 | 1 | 1215 | 1 | 1476 |
| QTJ33236.1 | 0.0 | 1 | 1214 | 1 | 1391 |
| ALB16165.1 | 1.94e-48 | 1 | 1032 | 1 | 1045 |
| ASA81954.1 | 1.94e-48 | 1 | 1032 | 1 | 1045 |
| ASA84007.1 | 1.94e-48 | 1 | 1032 | 1 | 1045 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4Q2W_A | 2.31e-19 | 423 | 538 | 165 | 278 | CrystalStructure of pneumococcal peptidoglycan hydrolase LytB [Streptococcus pneumoniae TIGR4] |
| 4PI7_A | 7.38e-17 | 355 | 561 | 22 | 228 | ChainA, Autolysin E [Staphylococcus aureus subsp. aureus Mu50],4PI9_A Chain A, Autolysin E [Staphylococcus aureus subsp. aureus Mu50],4PIA_A Chain A, Autolysin E [Staphylococcus aureus subsp. aureus Mu50] |
| 4PI8_A | 4.57e-16 | 355 | 561 | 22 | 228 | ChainA, Autolysin E [Staphylococcus aureus subsp. aureus Mu50] |
| 6FXO_A | 1.28e-14 | 406 | 541 | 71 | 226 | ChainA, Bifunctional autolysin [Staphylococcus aureus subsp. aureus Mu50] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P59205 | 5.69e-18 | 423 | 539 | 533 | 647 | Putative endo-beta-N-acetylglucosaminidase OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=lytB PE=1 SV=1 |
| P59206 | 6.24e-18 | 423 | 539 | 577 | 691 | Putative endo-beta-N-acetylglucosaminidase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=lytB PE=1 SV=1 |
| Q5HH31 | 1.15e-12 | 364 | 541 | 1049 | 1238 | Bifunctional autolysin OS=Staphylococcus aureus (strain COL) OX=93062 GN=atl PE=3 SV=1 |
| Q99V41 | 1.50e-12 | 406 | 541 | 1075 | 1230 | Bifunctional autolysin OS=Staphylococcus aureus (strain N315) OX=158879 GN=atl PE=1 SV=1 |
| Q931U5 | 1.50e-12 | 406 | 541 | 1075 | 1230 | Bifunctional autolysin OS=Staphylococcus aureus (strain Mu50 / ATCC 700699) OX=158878 GN=atl PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
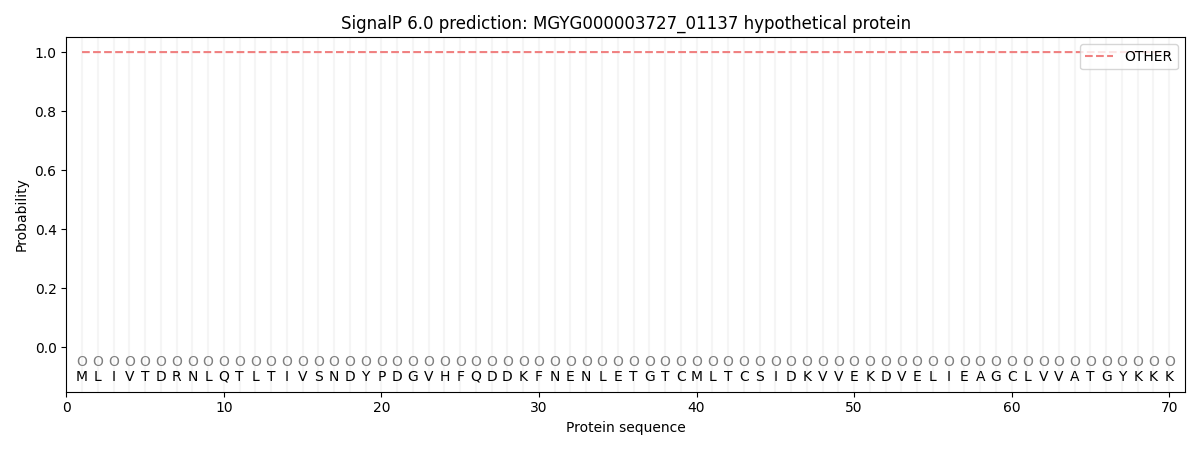
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000042 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
