You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003765_00051
You are here: Home > Sequence: MGYG000003765_00051
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella bergensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella bergensis | |||||||||||
| CAZyme ID | MGYG000003765_00051 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21904; End: 23445 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16144 | ARS_like | 1.32e-166 | 27 | 495 | 1 | 418 | uncharacterized arylsulfatase subfamily. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
| cd16146 | ARS_like | 1.39e-85 | 27 | 495 | 1 | 401 | uncharacterized arylsulfatase. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
| cd16145 | ARS_like | 1.05e-73 | 27 | 485 | 1 | 415 | uncharacterized arylsulfatase subfamily. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
| cd16027 | SGSH | 1.85e-72 | 27 | 500 | 1 | 373 | N-sulfoglucosamine sulfohydrolase (SGSH; sulfamidase). N-sulfoglucosamine sulfohydrolase (SGSH) belongs to the sulfatase family and catalyses the cleavage of N-linked sulfate groups from the GAGs heparin sulfate and heparin. The active site is characterized by the amino-acid sequence motif C(X)PSR that is highly conserved among most sulfatases. The cysteine residue is post-translationally converted to a formylglycine (FGly) residue, which is crucial for the catalytic process. Loss of function of SGSH results a disease called mucopolysaccharidosis type IIIA (Sanfilippo A syndrome), a fatal childhood-onset neurodegenerative disease with mild facial, visceral and skeletal abnormalities. |
| cd16025 | PAS_like | 1.44e-71 | 25 | 480 | 1 | 402 | Bacterial Arylsulfatase of Pseudomonas aeruginosa and related proteins. Sulfatases catalyze the hydrolysis of sulfate esters from wide range of substrates, including steroids, carbohydrates and proteins. Sulfate esters may be formed from various alcohols and amines. The biological roles of sulfatase includes the cycling of sulfur in the environment, in the degradation of sulfated glycosaminoglycans and glycolipids in the lysosome, and in remodeling sulfated glycosaminoglycans in the extracellular space. The sulfatases are essential for human metabolism. At least eight human monogenic diseases are caused by the deficiency of individual sulfatases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| EAR02039.1 | 3.20e-145 | 26 | 511 | 28 | 500 |
| QUT89376.1 | 1.01e-136 | 8 | 511 | 5 | 489 |
| ALJ59588.1 | 3.14e-135 | 8 | 511 | 5 | 489 |
| AKJ65363.1 | 5.55e-63 | 1 | 495 | 1 | 440 |
| QEF98196.1 | 7.55e-35 | 10 | 491 | 20 | 440 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6USS_A | 2.07e-160 | 1 | 508 | 1 | 503 | ChainA, Sulfatase [Bacteroides fragilis CAG:558],6USS_B Chain B, Sulfatase [Bacteroides fragilis CAG:558] |
| 6UST_A | 1.96e-61 | 26 | 492 | 4 | 446 | ChainA, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi],6UST_B Chain B, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi],6UST_C Chain C, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi],6UST_D Chain D, N-acetylgalactosamine 6-sulfate sulfatase [Hungatella hathewayi] |
| 7STT_A | 5.99e-60 | 22 | 507 | 2 | 441 | ChainA, N-acetylgalactosamine-6-sulfatase [Pedobacter yulinensis],7STU_A Chain A, N-acetylgalactosamine-6-sulfatase [Pedobacter yulinensis],7STV_A Chain A, N-acetylgalactosamine-6-sulfatase [Pedobacter yulinensis] |
| 6B0K_A | 4.89e-36 | 26 | 481 | 2 | 401 | Crystalstructure of Ps i-CgsB C78S in complex with k-carrapentaose [Pseudoalteromonas],6B0K_B Crystal structure of Ps i-CgsB C78S in complex with k-carrapentaose [Pseudoalteromonas],6B0K_C Crystal structure of Ps i-CgsB C78S in complex with k-carrapentaose [Pseudoalteromonas] |
| 6B0J_A | 4.97e-36 | 26 | 481 | 2 | 401 | Crystalstructure of Ps i-CgsB in complex with k-i-k-neocarrahexaose [Pseudoalteromonas],6B0J_B Crystal structure of Ps i-CgsB in complex with k-i-k-neocarrahexaose [Pseudoalteromonas],6B0J_C Crystal structure of Ps i-CgsB in complex with k-i-k-neocarrahexaose [Pseudoalteromonas],6B1V_A Crystal structure of Ps i-CgsB C78S in complex with i-neocarratetraose [Pseudoalteromonas],6B1V_B Crystal structure of Ps i-CgsB C78S in complex with i-neocarratetraose [Pseudoalteromonas],6B1V_C Crystal structure of Ps i-CgsB C78S in complex with i-neocarratetraose [Pseudoalteromonas] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KPJ9 | 1.92e-40 | 8 | 485 | 13 | 504 | Sulfatase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22020 PE=3 SV=1 |
| Q9C0V7 | 1.65e-28 | 26 | 485 | 11 | 513 | Uncharacterized sulfatase PB10D8.02c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBPB10D8.02c PE=3 SV=1 |
| P25549 | 1.92e-26 | 20 | 412 | 79 | 435 | Putative sulfatase AslA OS=Escherichia coli (strain K12) OX=83333 GN=aslA PE=3 SV=2 |
| Q8WNQ7 | 1.37e-24 | 14 | 428 | 17 | 385 | N-acetylgalactosamine-6-sulfatase OS=Sus scrofa OX=9823 GN=GALNS PE=2 SV=1 |
| Q571E4 | 1.82e-24 | 14 | 397 | 15 | 354 | N-acetylgalactosamine-6-sulfatase OS=Mus musculus OX=10090 GN=Galns PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
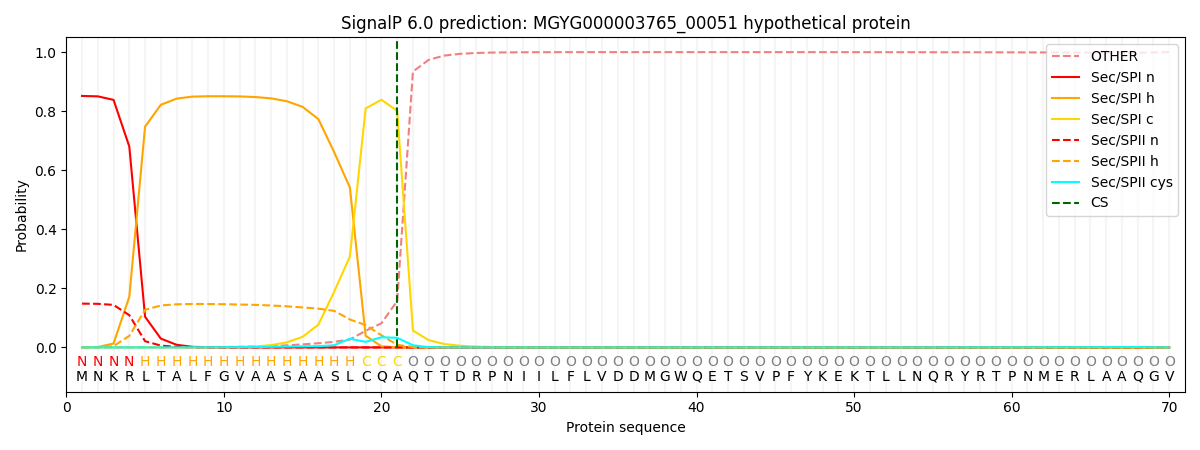
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000971 | 0.842373 | 0.155473 | 0.000585 | 0.000319 | 0.000245 |
