You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003772_00609
You are here: Home > Sequence: MGYG000003772_00609
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_C; Negativicutes; Veillonellales; Negativicoccaceae; ; | |||||||||||
| CAZyme ID | MGYG000003772_00609 | |||||||||||
| CAZy Family | GT28 | |||||||||||
| CAZyme Description | Processive diacylglycerol beta-glucosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4173; End: 5258 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT28 | 200 | 324 | 1.9e-28 | 0.7834394904458599 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd17507 | GT28_Beta-DGS-like | 8.17e-70 | 8 | 334 | 1 | 333 | beta-diglucosyldiacylglycerol synthase and similar proteins. beta-diglucosyldiacylglycerol synthase (processive diacylglycerol beta-glucosyltransferase EC 2.4.1.315) is involved in the biosynthesis of both the bilayer- and non-bilayer-forming membrane glucolipids. This family of glycosyltransferases also contains plant major galactolipid synthase (chloroplastic monogalactosyldiacylglycerol synthase 1 EC 2.4.1.46). Glycosyltransferases catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. The acceptor molecule can be a lipid, a protein, a heterocyclic compound, or another carbohydrate residue. The structures of the formed glycoconjugates are extremely diverse, reflecting a wide range of biological functions. The members of this family share a common GTB topology, one of the two protein topologies observed for nucleotide-sugar-dependent glycosyltransferases. GTB proteins have distinct N- and C- terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| PRK13609 | PRK13609 | 6.55e-55 | 3 | 322 | 2 | 326 | diacylglycerol glucosyltransferase; Provisional |
| COG0707 | MurG | 4.17e-40 | 10 | 354 | 4 | 351 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
| PRK13608 | PRK13608 | 2.52e-29 | 8 | 316 | 8 | 320 | diacylglycerol glucosyltransferase; Provisional |
| PLN02605 | PLN02605 | 3.56e-24 | 8 | 308 | 1 | 321 | monogalactosyldiacylglycerol synthase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBK24795.1 | 3.64e-106 | 10 | 334 | 10 | 336 |
| AOH38912.1 | 5.89e-105 | 10 | 358 | 9 | 364 |
| BBK25634.1 | 1.02e-104 | 10 | 358 | 9 | 364 |
| AVO75296.1 | 1.37e-101 | 5 | 337 | 3 | 340 |
| CCC74010.1 | 1.37e-101 | 5 | 337 | 3 | 340 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B9J2U2 | 9.25e-43 | 3 | 326 | 2 | 330 | Processive diacylglycerol beta-glucosyltransferase OS=Bacillus cereus (strain Q1) OX=361100 GN=ugtP PE=3 SV=1 |
| B7HU46 | 9.25e-43 | 3 | 326 | 2 | 330 | Processive diacylglycerol beta-glucosyltransferase OS=Bacillus cereus (strain AH187) OX=405534 GN=ugtP PE=3 SV=1 |
| Q73DZ5 | 9.25e-43 | 3 | 326 | 2 | 330 | Processive diacylglycerol beta-glucosyltransferase OS=Bacillus cereus (strain ATCC 10987 / NRS 248) OX=222523 GN=ugtP PE=3 SV=1 |
| C1EWE6 | 1.29e-42 | 3 | 326 | 2 | 330 | Processive diacylglycerol beta-glucosyltransferase OS=Bacillus cereus (strain 03BB102) OX=572264 GN=ugtP PE=3 SV=1 |
| Q63GD0 | 1.80e-42 | 3 | 326 | 2 | 330 | Processive diacylglycerol beta-glucosyltransferase OS=Bacillus cereus (strain ZK / E33L) OX=288681 GN=ugtP PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
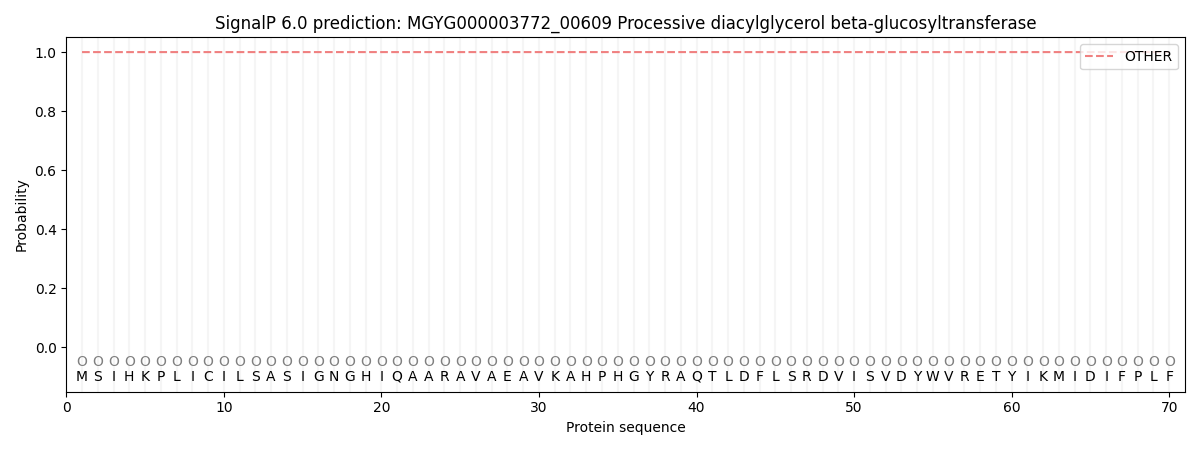
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000035 | 0.000009 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
