You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003777_00815
Basic Information
help
| Species |
|
| Lineage |
Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Muricomes;
|
| CAZyme ID |
MGYG000003777_00815
|
| CAZy Family |
GH106 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000003777 |
2405889 |
MAG |
Canada |
North America |
|
| Gene Location |
Start: 12186;
End: 15251
Strand: -
|
No EC number prediction in MGYG000003777_00815.
| Family |
Start |
End |
Evalue |
family coverage |
| GH106 |
8 |
848 |
4e-74 |
0.8434466019417476 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam17132
|
Glyco_hydro_106 |
2.17e-14 |
213 |
764 |
349 |
870 |
alpha-L-rhamnosidase. |
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
| 5MWK_A
|
6.10e-13 |
190 |
981 |
339 |
1061 |
Glycosidehydrolase BT_0986 [Bacteroides thetaiotaomicron] |
| 5MQM_A
|
1.38e-12 |
190 |
981 |
339 |
1061 |
Glycosidehydrolase BT_0986 [Bacteroides thetaiotaomicron],5MQN_A Glycoside hydrolase BT_0986 [Bacteroides thetaiotaomicron] |
| 6Q2F_A
|
7.25e-10 |
212 |
1013 |
407 |
1133 |
Structureof Rhamnosidase from Novosphingobium sp. PP1Y [Novosphingobium sp. PP1Y] |
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
|
T2KNA8
|
5.95e-10 |
113 |
982 |
112 |
1096 |
Putative beta-glucuronidase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_22040 PE=2 SV=1 |
This protein is predicted as OTHER
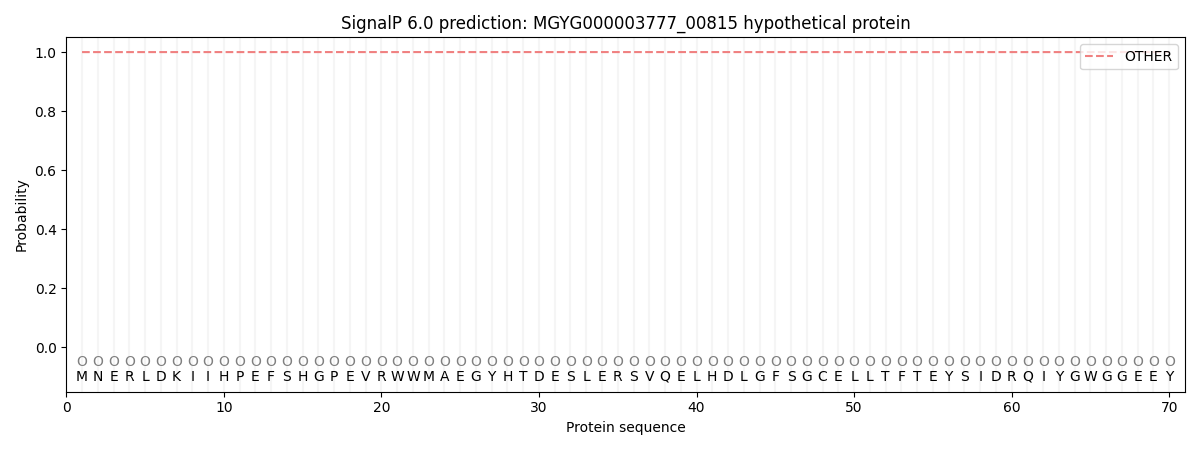
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000063
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000003777_00815.

