You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003867_00641
You are here: Home > Sequence: MGYG000003867_00641
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Catenibacillus; | |||||||||||
| CAZyme ID | MGYG000003867_00641 | |||||||||||
| CAZy Family | GH9 | |||||||||||
| CAZyme Description | Cellodextrinase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 6148; End: 7239 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH9 | 6 | 351 | 1.2e-81 | 0.80622009569378 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00759 | Glyco_hydro_9 | 6.73e-50 | 5 | 350 | 84 | 374 | Glycosyl hydrolase family 9. |
| PLN00119 | PLN00119 | 5.73e-08 | 76 | 306 | 189 | 458 | endoglucanase |
| PLN02909 | PLN02909 | 1.32e-07 | 76 | 284 | 192 | 409 | Endoglucanase |
| PLN03009 | PLN03009 | 1.40e-07 | 75 | 356 | 187 | 492 | cellulase |
| PLN02171 | PLN02171 | 2.40e-06 | 128 | 300 | 239 | 433 | endoglucanase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADD61951.1 | 3.88e-172 | 1 | 354 | 195 | 555 |
| AGS53175.1 | 1.60e-109 | 1 | 355 | 176 | 529 |
| AMO13180.1 | 9.62e-107 | 1 | 355 | 192 | 546 |
| ABW39324.1 | 2.35e-106 | 1 | 355 | 189 | 543 |
| ABW39322.1 | 7.64e-102 | 1 | 355 | 189 | 543 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3X17_A | 9.15e-68 | 1 | 351 | 210 | 554 | Crystalstructure of metagenome-derived glycoside hydrolase family 9 endoglucanase [uncultured bacterium],3X17_B Crystal structure of metagenome-derived glycoside hydrolase family 9 endoglucanase [uncultured bacterium] |
| 6DHT_A | 6.47e-66 | 1 | 354 | 202 | 565 | Bacteroidesovatus GH9 Bacova_02649 [Bacteroides ovatus ATCC 8483] |
| 4CJ0_A | 7.33e-65 | 1 | 354 | 208 | 549 | ChainA, ENDOGLUCANASE D [Acetivibrio thermocellus],4CJ1_A Chain A, ENDOGLUCANASE D [Acetivibrio thermocellus] |
| 1CLC_A | 9.42e-65 | 1 | 354 | 222 | 563 | ChainA, ENDOGLUCANASE CELD; EC: 3.2.1.4 [Acetivibrio thermocellus] |
| 3EZ8_A | 5.87e-62 | 10 | 353 | 187 | 531 | CrystalStructure of endoglucanase Cel9A from the thermoacidophilic Alicyclobacillus acidocaldarius [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3GZK_A Structure of A. Acidocaldarius Cellulase CelA [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3H2W_A Structure of A. acidocaldarius cellulase CelA in complex with cellobiose [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3H3K_A Structure of A. acidocaldarius cellulase CelA in complex with cellotetraose [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3RX5_A structure of AaCel9A in complex with cellotriose-like isofagomine [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3RX7_A Structure of AaCel9A in complex with cellotetraose-like isofagomine [Alicyclobacillus acidocaldarius subsp. acidocaldarius],3RX8_A structure of AaCel9A in complex with cellobiose-like isofagomine [Alicyclobacillus acidocaldarius subsp. acidocaldarius] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P23658 | 3.87e-94 | 5 | 348 | 186 | 540 | Cellodextrinase OS=Butyrivibrio fibrisolvens OX=831 GN=ced1 PE=1 SV=1 |
| A7LXT3 | 5.09e-65 | 1 | 354 | 216 | 579 | Xyloglucan-specific endo-beta-1,4-glucanase BoGH9A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02649 PE=1 SV=1 |
| P0C2S4 | 4.01e-64 | 1 | 354 | 208 | 549 | Endoglucanase D (Fragment) OS=Acetivibrio thermocellus OX=1515 GN=celD PE=1 SV=1 |
| A3DDN1 | 6.15e-64 | 1 | 354 | 232 | 573 | Endoglucanase D OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celD PE=1 SV=1 |
| A3DCH1 | 9.04e-56 | 1 | 359 | 424 | 817 | Cellulose 1,4-beta-cellobiosidase OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celK PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
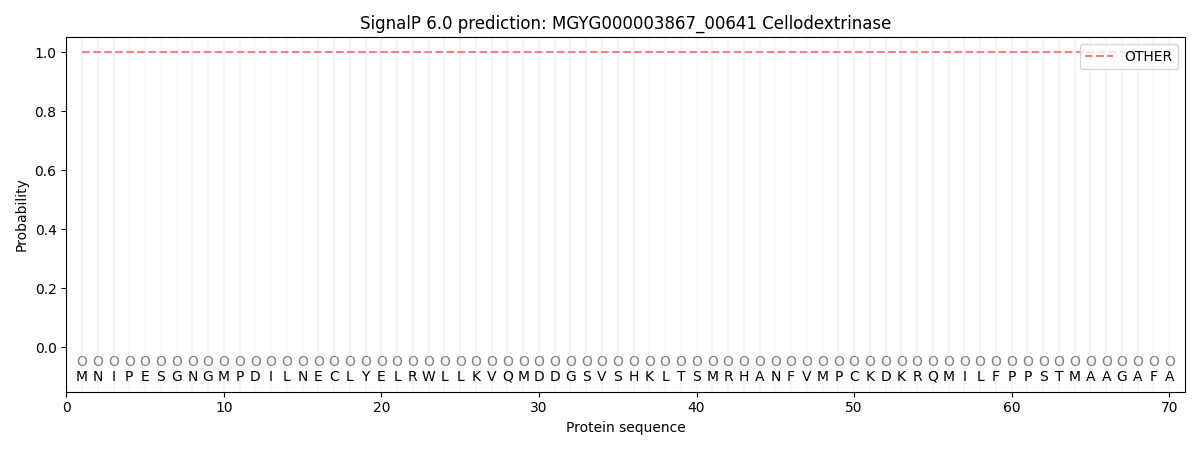
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999990 | 0.000048 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
