You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003922_00995
You are here: Home > Sequence: MGYG000003922_00995
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp014385165 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp014385165 | |||||||||||
| CAZyme ID | MGYG000003922_00995 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 89798; End: 91732 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06542 | GH18_EndoS-like | 1.11e-60 | 189 | 457 | 1 | 255 | Endo-beta-N-acetylglucosaminidases are bacterial chitinases that hydrolyze the chitin core of various asparagine (N)-linked glycans and glycoproteins. The endo-beta-N-acetylglucosaminidases have a glycosyl hydrolase family 18 (GH18) catalytic domain. Some members also have an additional C-terminal glycosyl hydrolase family 20 (GH20) domain while others have an N-terminal domain of unknown function (pfam08522). Members of this family include endo-beta-N-acetylglucosaminidase S (EndoS) from Streptococcus pyogenes, EndoF1, EndoF2, EndoF3, and EndoH from Flavobacterium meningosepticum, and EndoE from Enterococcus faecalis. EndoS is a secreted endoglycosidase from Streptococcus pyogenes that specifically hydrolyzes the glycan on human IgG between two core N-acetylglucosamine residues. EndoE is a secreted endoglycosidase, encoded by the ndoE gene in Enterococcus faecalis, that hydrolyzes the glycan on human RNase B. |
| pfam08522 | DUF1735 | 1.93e-16 | 45 | 161 | 2 | 118 | Domain of unknown function (DUF1735). This domain of unknown function is found in a number of bacterial proteins including acylhydrolases. The structure of this domain has a beta-sandwich fold. |
| pfam00704 | Glyco_hydro_18 | 1.65e-09 | 223 | 381 | 26 | 176 | Glycosyl hydrolases family 18. |
| cd02871 | GH18_chitinase_D-like | 8.24e-06 | 218 | 321 | 28 | 129 | GH18 domain of Chitinase D (ChiD). ChiD, a chitinase found in Bacillus circulans, hydrolyzes the 1,4-beta-linkages of N-acetylglucosamine in chitin and chitodextrins. The domain architecture of ChiD includes a catalytic glycosyl hydrolase family 18 (GH18) domain, a chitin-binding domain, and a fibronectin type III domain. The chitin-binding and fibronectin type III domains are located either N-terminal or C-terminal to the catalytic domain. This family includes exochitinase Chi36 from Bacillus cereus. |
| cd02878 | GH18_zymocin_alpha | 2.19e-05 | 247 | 380 | 45 | 185 | Zymocin, alpha subunit. Zymocin is a heterotrimeric enzyme that inhibits yeast cell cycle progression. The zymocin alpha subunit has a chitinase activity that is essential for holoenzyme action from the cell exterior while the gamma subunit contains the intracellular toxin responsible for G1 phase cell cycle arrest. The zymocin alpha and beta subunits are thought to act from the cell's exterior by docking to the cell wall-associated chitin, thus mediating gamma-toxin translocation. The alpha subunit has an eight-stranded TIM barrel fold similar to that of family 18 glycosyl hydrolases such as hevamine, chitolectin, and chitobiase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDO69363.1 | 0.0 | 1 | 644 | 4 | 660 |
| QUT92946.1 | 0.0 | 1 | 643 | 4 | 657 |
| QPH59421.1 | 4.57e-268 | 1 | 643 | 1 | 658 |
| QMI80605.1 | 6.48e-268 | 1 | 643 | 1 | 658 |
| QBJ19120.1 | 6.48e-268 | 1 | 643 | 1 | 658 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6T8I_A | 3.71e-58 | 78 | 466 | 52 | 429 | Crystalstructure of wild type EndoBT-3987 from Bacteroides thetaiotamicron VPI-5482 [Bacteroides thetaiotaomicron VPI-5482],6T8K_A Crystal structure of Bacteroides thetaiotamicron EndoBT-3987 in complex with Man9GlcNAc product in P1 [Bacteroides thetaiotaomicron VPI-5482],6T8K_B Crystal structure of Bacteroides thetaiotamicron EndoBT-3987 in complex with Man9GlcNAc product in P1 [Bacteroides thetaiotaomicron VPI-5482],6T8L_A Crystal structure of Bacteroides thetaiotamicron EndoBT-3987 with Man9GlcNAc product in P212121 [Bacteroides thetaiotaomicron VPI-5482],6TCW_A Crystal structure of Bacteroides thetaiotamicron EndoBT-3987 with Man5GlcNAc product [Bacteroides thetaiotaomicron VPI-5482],7NWF_A Chain A, Endo-beta-N-acetylglucosaminidase F1 [Bacteroides thetaiotaomicron VPI-5482] |
| 3POH_A | 2.63e-56 | 78 | 466 | 52 | 429 | Crystalstructure of an endo-beta-N-acetylglucosaminidase (BT_3987) from BACTEROIDES THETAIOTAOMICRON VPI-5482 at 1.55 A resolution [Bacteroides thetaiotaomicron VPI-5482] |
| 6TCV_B | 7.01e-56 | 78 | 466 | 52 | 429 | Crystalstructure of Bacteroides thetaiotamicron EndoBT-3987 in complex with Man9GlcNAc2Asn substrate [Bacteroides thetaiotaomicron VPI-5482] |
| 2EBN_A | 1.05e-49 | 180 | 471 | 1 | 288 | CRYSTALSTRUCTURE OF ENDO-BETA-N-ACETYLGLUCOSAMINIDASE F1, AN ALPHA(SLASH)BETA-BARREL ENZYME ADAPTED FOR A COMPLEX SUBSTRATE [Elizabethkingia meningoseptica] |
| 1EDT_A | 2.64e-42 | 182 | 437 | 2 | 245 | ChainA, ENDO-BETA-N-ACETYLGLUCOSAMINIDASE H, ENDO H [Streptomyces plicatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P36911 | 2.19e-48 | 180 | 471 | 51 | 338 | Endo-beta-N-acetylglucosaminidase F1 OS=Elizabethkingia meningoseptica OX=238 GN=endOF1 PE=1 SV=1 |
| P04067 | 8.11e-41 | 191 | 437 | 53 | 287 | Endo-beta-N-acetylglucosaminidase H OS=Streptomyces plicatus OX=1922 PE=1 SV=1 |
| P80036 | 4.47e-37 | 186 | 457 | 52 | 307 | Endo-beta-N-acetylglucosaminidase OS=Flavobacterium sp. (strain SK1022) OX=148444 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
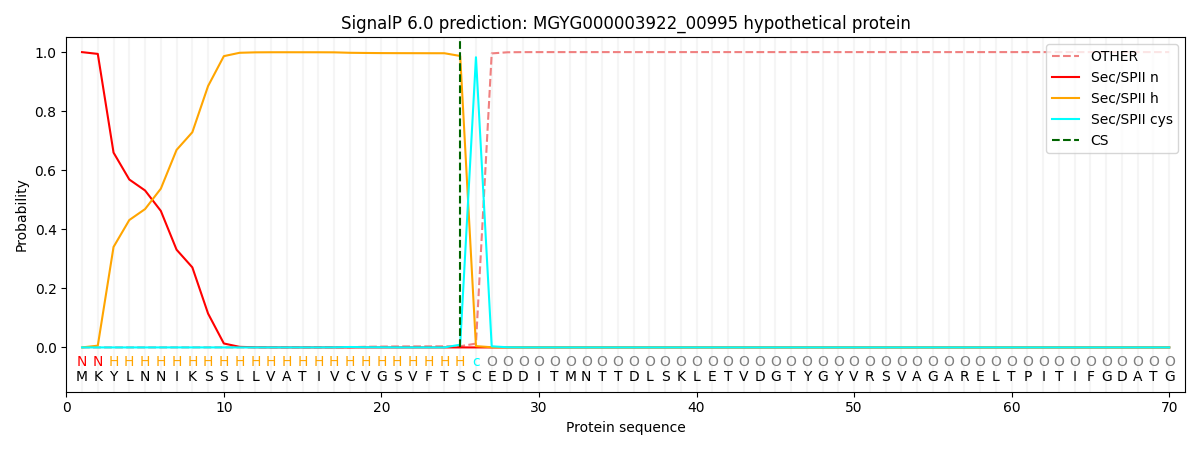
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000077 | 0.999947 | 0.000000 | 0.000000 | 0.000000 |
