You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003941_00192
You are here: Home > Sequence: MGYG000003941_00192
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; | |||||||||||
| CAZyme ID | MGYG000003941_00192 | |||||||||||
| CAZy Family | GH38 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 53387; End: 56008 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH38 | 75 | 220 | 5.7e-29 | 0.5278810408921933 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0383 | AMS1 | 2.24e-18 | 135 | 873 | 286 | 943 | Alpha-mannosidase [Carbohydrate transport and metabolism]. |
| pfam07748 | Glyco_hydro_38C | 5.04e-14 | 513 | 711 | 1 | 180 | Glycosyl hydrolases family 38 C-terminal domain. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. |
| cd10789 | GH38N_AMII_ER_cytosolic | 1.21e-10 | 105 | 217 | 60 | 169 | N-terminal catalytic domain of endoplasmic reticulum(ER)/cytosolic class II alpha-mannosidases; glycoside hydrolase family 38 (GH38). The subfamily is represented by Saccharomyces cerevisiae vacuolar alpha-mannosidase Ams1, rat ER/cytosolic alpha-mannosidase Man2C1, and similar proteins. Members in this family share high sequence similarity. None of them have any classical signal sequence or membrane spanning domains, which are typical of sorting or targeting signals. Ams1 functions as a second resident vacuolar hydrolase in S. cerevisiae. It aids in recycling macromolecular components of the cell through hydrolysis of terminal, non-reducing alpha-d-mannose residues. Ams1 utilizes both the cytoplasm to vacuole targeting (Cvt, nutrient-rich conditions) and autophagic (starvation conditions) pathways for biosynthetic delivery to the vacuole. Man2C1is involved in oligosaccharide catabolism in both the ER and cytosol. It can catalyze the cobalt-dependent cleavage of alpha 1,2-, alpha 1,3-, and alpha 1,6-linked mannose residues. Members in this family are retaining glycosyl hydrolases of family GH38 that employs a two-step mechanism involving the formation of a covalent glycosyl-enzyme complex. Two carboxylic acids positioned within the active site act in concert: one as a catalytic nucleophile and the other as a general acid/base catalyst. |
| pfam01074 | Glyco_hydro_38 | 1.32e-10 | 105 | 230 | 60 | 190 | Glycosyl hydrolases family 38 N-terminal domain. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. |
| cd10786 | GH38N_AMII_like | 7.23e-09 | 61 | 219 | 19 | 172 | N-terminal catalytic domain of class II alpha-mannosidases and similar proteins; glycoside hydrolase family 38 (GH38). Alpha-mannosidases (EC 3.2.1.24) are extensively found in eukaryotes and play important roles in the processing of newly formed N-glycans and in degradation of mature glycoproteins. A deficiency of this enzyme causes the lysosomal storage disease alpha-mannosidosis. Many bacterial and archaeal species also possess putative alpha-mannosidases, but their activity and specificity is largely unknown. Based on different functional characteristics and sequence homology, alpha-mannosidases have been organized into two classes (class I, belonging to glycoside hydrolase family 47, and class II, belonging to glycoside hydrolase family 38). Members of this family corresponds to class II alpha-mannosidases (alphaMII), which contain intermediate Golgi alpha-mannosidases II, acidic lysosomal alpha-mannosidases, animal sperm and epididymal alpha -mannosidases, neutral ER/cytosolic alpha-mannosidases, and some putative prokaryotic alpha-mannosidases. AlphaMII possess a-1,3, a-1,6, and a-1,2 hydrolytic activity, and catalyzes the degradation of N-linked oligosaccharides. The N-terminal catalytic domain of alphaMII adopts a structure consisting of parallel 7-stranded beta/alpha barrel. Members in this family are retaining glycosyl hydrolases of family GH38 that employs a two-step mechanism involving the formation of a covalent glycosyl enzyme complex. Two carboxylic acids positioned within the active site act in concert: one as a catalytic nucleophile and the other as a general acid/base catalyst. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBL06663.1 | 0.0 | 1 | 873 | 1 | 880 |
| BBL11622.1 | 0.0 | 39 | 873 | 43 | 883 |
| BBL08830.1 | 0.0 | 39 | 873 | 43 | 883 |
| BBL00893.1 | 0.0 | 4 | 873 | 11 | 882 |
| QNL41054.1 | 0.0 | 35 | 871 | 6 | 852 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
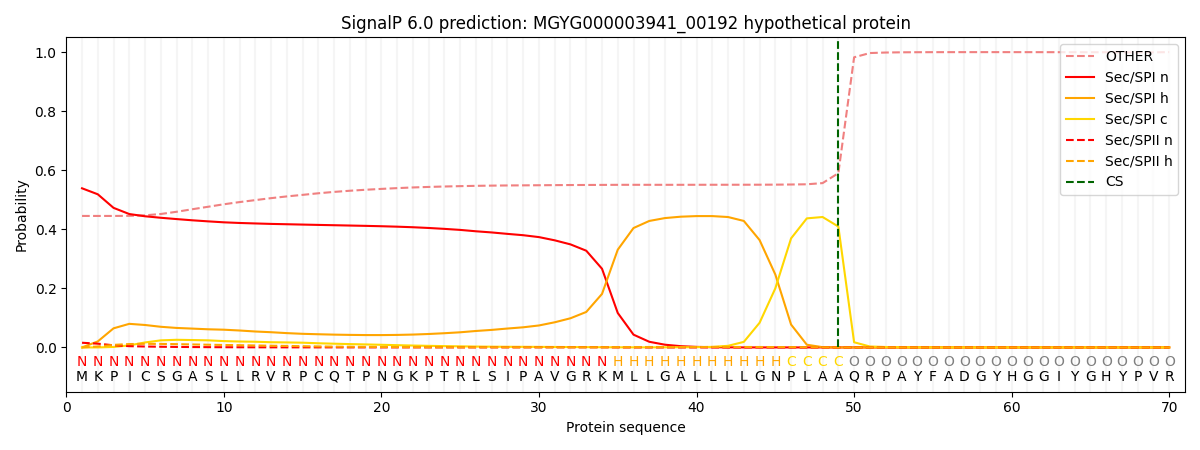
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.451457 | 0.529250 | 0.018326 | 0.000449 | 0.000283 | 0.000233 |
