You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003952_01179
You are here: Home > Sequence: MGYG000003952_01179
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp900021155 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp900021155 | |||||||||||
| CAZyme ID | MGYG000003952_01179 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 39570; End: 41081 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 47 | 402 | 2.2e-61 | 0.9230769230769231 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.92e-48 | 20 | 332 | 81 | 396 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| PLN02188 | PLN02188 | 3.27e-14 | 3 | 328 | 6 | 345 | polygalacturonase/glycoside hydrolase family protein |
| pfam00295 | Glyco_hydro_28 | 3.92e-12 | 172 | 330 | 88 | 237 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN03003 | PLN03003 | 2.34e-11 | 2 | 296 | 4 | 284 | Probable polygalacturonase At3g15720 |
| pfam12708 | Pectate_lyase_3 | 2.40e-10 | 23 | 310 | 3 | 213 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AMJ66331.1 | 1.63e-114 | 1 | 484 | 1 | 489 |
| AMR25945.1 | 4.80e-112 | 19 | 483 | 19 | 489 |
| AWM31556.1 | 1.40e-102 | 10 | 485 | 13 | 496 |
| QHW33703.1 | 2.86e-77 | 22 | 473 | 5 | 457 |
| QJD86476.1 | 6.13e-75 | 20 | 469 | 9 | 456 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OLP_A | 2.90e-23 | 22 | 246 | 45 | 275 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 3JUR_A | 3.19e-21 | 27 | 246 | 33 | 265 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 4MXN_A | 1.73e-15 | 22 | 248 | 22 | 225 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
| 2UVE_A | 2.46e-11 | 17 | 251 | 152 | 413 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P35339 | 1.17e-10 | 22 | 246 | 41 | 238 | Exopolygalacturonase OS=Zea mays OX=4577 GN=PG2C PE=2 SV=1 |
| P15922 | 5.42e-10 | 23 | 251 | 153 | 406 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
| Q05967 | 3.37e-09 | 17 | 284 | 22 | 271 | Polygalacturonase OS=Nicotiana tabacum OX=4097 GN=PG1 PE=2 SV=1 |
| Q9LW07 | 2.88e-08 | 23 | 246 | 25 | 215 | Probable polygalacturonase At3g15720 OS=Arabidopsis thaliana OX=3702 GN=At3g15720 PE=3 SV=1 |
| Q7M1E7 | 7.40e-08 | 19 | 246 | 56 | 260 | Polygalacturonase OS=Chamaecyparis obtusa OX=13415 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
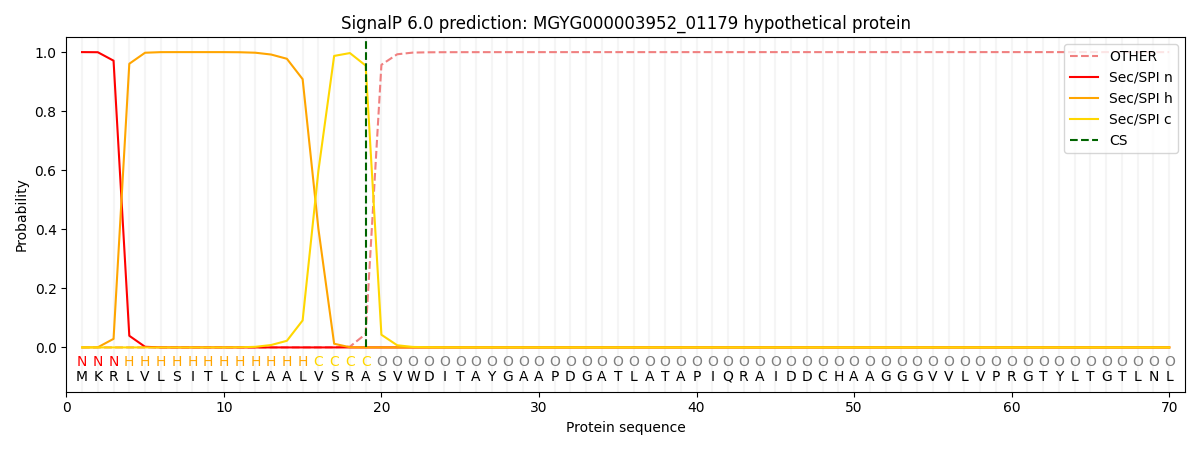
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000435 | 0.998648 | 0.000248 | 0.000214 | 0.000209 | 0.000201 |
