You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003989_00189
You are here: Home > Sequence: MGYG000003989_00189
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes sp002161445 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes sp002161445 | |||||||||||
| CAZyme ID | MGYG000003989_00189 | |||||||||||
| CAZy Family | GT90 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 5583; End: 6560 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam05686 | Glyco_transf_90 | 6.75e-11 | 206 | 300 | 207 | 302 | Glycosyl transferase family 90. This family of glycosyl transferases are specifically (mannosyl) glucuronoxylomannan/galactoxylomannan -beta 1,2-xylosyltransferases, EC:2.4.2.-. |
| smart00672 | CAP10 | 8.99e-11 | 206 | 299 | 137 | 231 | Putative lipopolysaccharide-modifying enzyme. |
| pfam13524 | Glyco_trans_1_2 | 0.002 | 243 | 295 | 30 | 76 | Glycosyl transferases group 1. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQS97119.1 | 4.16e-105 | 7 | 323 | 8 | 319 |
| QQT34018.1 | 4.16e-105 | 7 | 323 | 8 | 319 |
| QQT24222.1 | 6.29e-102 | 7 | 323 | 8 | 319 |
| QSG83178.1 | 9.27e-94 | 11 | 324 | 11 | 321 |
| QYS92068.1 | 1.31e-93 | 11 | 324 | 11 | 321 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39907 | 3.86e-62 | 52 | 318 | 51 | 312 | O-glucosyltransferase LpsA OS=Dichelobacter nodosus OX=870 GN=lpsA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
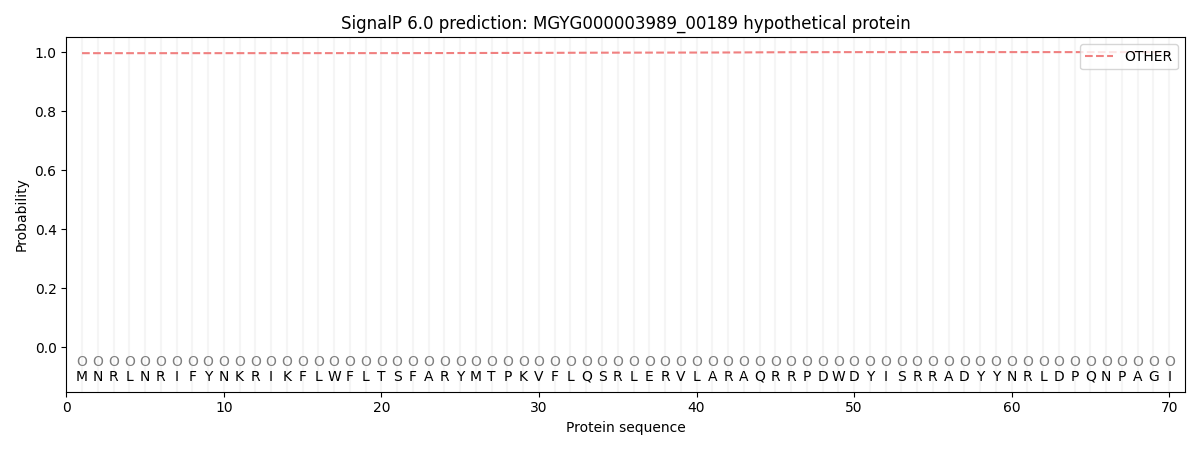
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.996662 | 0.002924 | 0.000371 | 0.000018 | 0.000009 | 0.000024 |
