You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000003994_01302
You are here: Home > Sequence: MGYG000003994_01302
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA11471; UBA11471; | |||||||||||
| CAZyme ID | MGYG000003994_01302 | |||||||||||
| CAZy Family | PL10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9767; End: 10900 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL10 | 62 | 350 | 2.1e-120 | 0.9930555555555556 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam09492 | Pec_lyase | 1.11e-146 | 62 | 352 | 1 | 289 | Pectic acid lyase. Members of this family are isozymes of pectate lyase (EC:4.2.2.2), also called polygalacturonic transeliminase and alpha-1,4-D-endopolygalacturonic acid lyase. |
| TIGR02474 | pec_lyase | 8.90e-138 | 62 | 352 | 1 | 290 | pectate lyase, PelA/Pel-15E family. Members of this family are isozymes of pectate lyase (EC 4.2.2.2), also called polygalacturonic transeliminase and alpha-1,4-D-endopolygalacturonic acid lyase. [Energy metabolism, Biosynthesis and degradation of polysaccharides] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDH52817.1 | 7.87e-161 | 1 | 361 | 1 | 361 |
| QRQ48738.1 | 1.29e-160 | 1 | 364 | 1 | 368 |
| QUT45489.1 | 1.29e-160 | 1 | 364 | 1 | 368 |
| QUT30912.1 | 6.42e-160 | 1 | 361 | 1 | 361 |
| SCV06361.1 | 2.60e-159 | 1 | 361 | 1 | 361 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1R76_A | 2.41e-70 | 47 | 360 | 78 | 406 | ChainA, pectate lyase [Niveispirillum irakense] |
| 1GXM_A | 3.57e-62 | 55 | 352 | 37 | 323 | Family10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus],1GXM_B Family 10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus],1GXN_A Family 10 polysaccharide lyase from Cellvibrio cellulosa [Cellvibrio japonicus] |
| 1GXO_A | 5.53e-61 | 55 | 352 | 37 | 323 | MutantD189A of Family 10 polysaccharide lyase from Cellvibrio cellulosa in complex with trigalaturonic acid [Cellvibrio japonicus] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
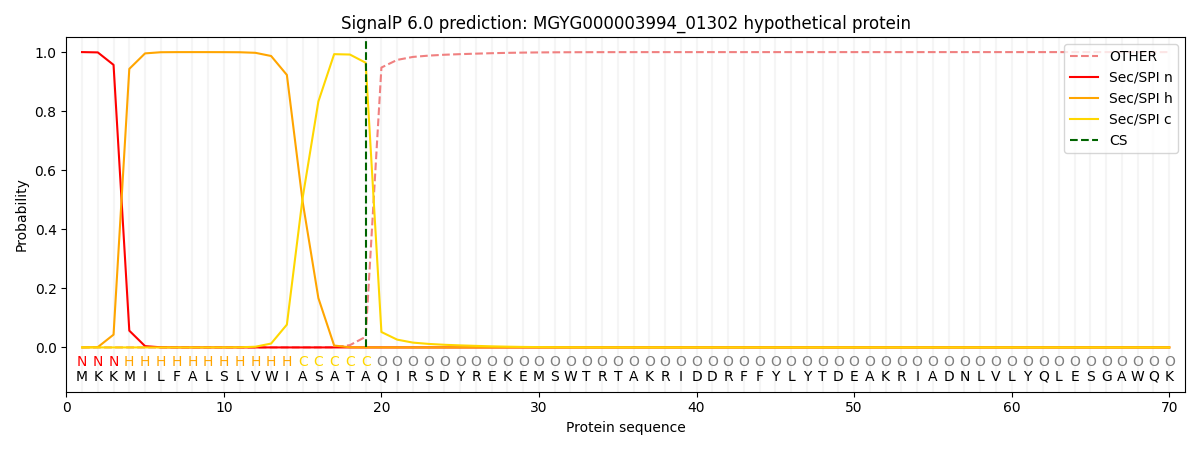
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000367 | 0.998978 | 0.000184 | 0.000165 | 0.000154 | 0.000153 |
