You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004017_02047
You are here: Home > Sequence: MGYG000004017_02047
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | HGM11588 sp900754755 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; HGM11588; HGM11588 sp900754755 | |||||||||||
| CAZyme ID | MGYG000004017_02047 | |||||||||||
| CAZy Family | GH109 | |||||||||||
| CAZyme Description | Alpha-N-acetylgalactosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 19577; End: 20824 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 8 | 400 | 4e-154 | 0.9874686716791979 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0673 | MviM | 3.12e-22 | 9 | 366 | 3 | 317 | Predicted dehydrogenase [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEF80535.1 | 1.40e-184 | 7 | 403 | 6 | 402 |
| ANY74959.1 | 8.93e-171 | 3 | 405 | 5 | 402 |
| AEE95391.1 | 7.31e-169 | 10 | 414 | 9 | 406 |
| QTH41090.1 | 5.20e-167 | 10 | 414 | 10 | 411 |
| QTH41319.1 | 2.59e-157 | 10 | 402 | 8 | 401 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2IXA_A | 1.23e-103 | 10 | 403 | 21 | 436 | A-zyme,N-acetylgalactosaminidase [Elizabethkingia meningoseptica],2IXB_A Crystal structure of N-ACETYLGALACTOSAMINIDASE in complex with GalNAC [Elizabethkingia meningoseptica] |
| 6T2B_A | 1.77e-81 | 10 | 400 | 43 | 440 | Glycosidehydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_B Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_C Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila],6T2B_D Glycoside hydrolase family 109 from Akkermansia muciniphila in complex with GalNAc and NAD+. [Akkermansia muciniphila] |
| 3EC7_A | 1.54e-09 | 7 | 164 | 21 | 176 | CrystalStructure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_B Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_C Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_D Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_E Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_F Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_G Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],3EC7_H Crystal Structure of Putative Dehydrogenase from Salmonella typhimurium LT2 [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 3MOI_A | 6.58e-06 | 10 | 157 | 3 | 145 | Thecrystal structure of the putative dehydrogenase from Bordetella bronchiseptica RB50 [Bordetella bronchiseptica] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B2FLK4 | 1.27e-105 | 7 | 398 | 31 | 441 | Glycosyl hydrolase family 109 protein OS=Stenotrophomonas maltophilia (strain K279a) OX=522373 GN=Smlt4431 PE=3 SV=1 |
| A4Q8G1 | 5.49e-105 | 4 | 399 | 47 | 460 | Alpha-N-acetylgalactosaminidase OS=Tannerella forsythia OX=28112 GN=nagA PE=3 SV=1 |
| A6LB54 | 3.28e-103 | 10 | 399 | 50 | 458 | Glycosyl hydrolase family 109 protein OS=Parabacteroides distasonis (strain ATCC 8503 / DSM 20701 / CIP 104284 / JCM 5825 / NCTC 11152) OX=435591 GN=BDI_1155 PE=3 SV=1 |
| A4Q8F7 | 6.74e-103 | 10 | 403 | 21 | 436 | Alpha-N-acetylgalactosaminidase OS=Elizabethkingia meningoseptica OX=238 GN=nagA PE=1 SV=1 |
| Q01S58 | 6.43e-96 | 9 | 400 | 42 | 434 | Glycosyl hydrolase family 109 protein OS=Solibacter usitatus (strain Ellin6076) OX=234267 GN=Acid_6590 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
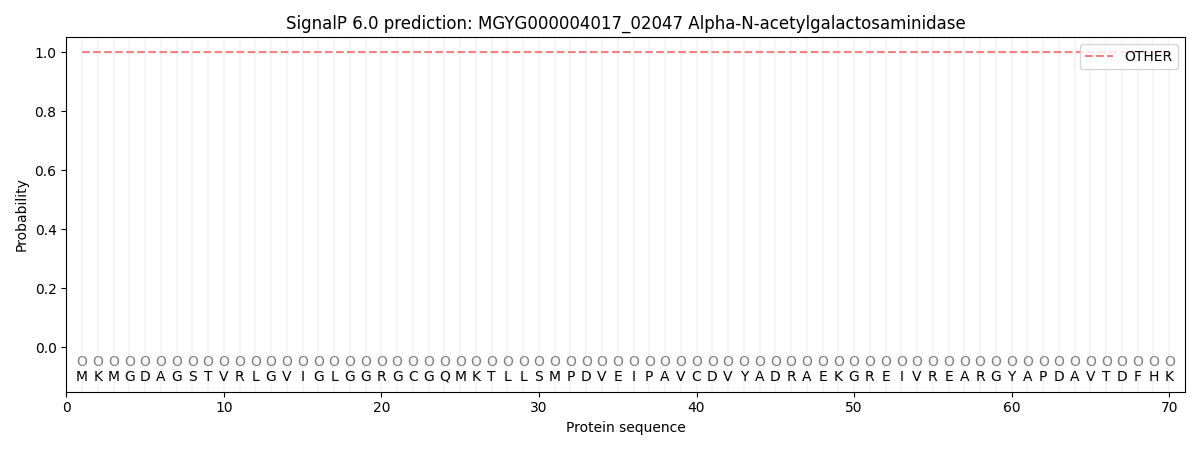
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000030 | 0.000002 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
