You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004021_00397
You are here: Home > Sequence: MGYG000004021_00397
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UBA11471 sp900542765 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; UBA11471; UBA11471; UBA11471 sp900542765 | |||||||||||
| CAZyme ID | MGYG000004021_00397 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 184511; End: 185932 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 164 | 331 | 3.1e-42 | 0.7821782178217822 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00656 | Amb_all | 2.69e-32 | 159 | 330 | 18 | 186 | Amb_all domain. |
| COG3866 | PelB | 5.15e-31 | 17 | 393 | 21 | 342 | Pectate lyase [Carbohydrate transport and metabolism]. |
| pfam00544 | Pec_lyase_C | 1.05e-16 | 176 | 330 | 52 | 211 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADE83376.1 | 1.31e-120 | 25 | 394 | 43 | 421 |
| QVJ81552.1 | 1.85e-120 | 25 | 394 | 43 | 421 |
| AAW84045.1 | 2.39e-112 | 65 | 398 | 1 | 328 |
| AHJ96631.1 | 1.83e-48 | 121 | 394 | 79 | 364 |
| AYN03314.1 | 1.68e-47 | 32 | 394 | 58 | 361 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1VBL_A | 1.41e-17 | 164 | 348 | 136 | 334 | Structureof the thermostable pectate lyase PL 47 [Bacillus sp. TS-47] |
| 3ZSC_A | 1.79e-17 | 176 | 306 | 83 | 213 | Catalyticfunction and substrate recognition of the pectate lyase from Thermotoga maritima [Thermotoga maritima] |
| 3KRG_A | 1.03e-12 | 184 | 307 | 150 | 296 | ChainA, Pectate lyase [Bacillus subtilis] |
| 5AMV_A | 4.33e-12 | 184 | 307 | 150 | 296 | Structuralinsights into the loss of catalytic competence in pectate lyase at low pH [Bacillus subtilis],5X2I_A Polygalacturonate Lyase by Fusing with a Self-assembling Amphipathic Peptide [Bacillus subtilis subsp. subtilis str. 168] |
| 1BN8_A | 4.71e-12 | 184 | 307 | 171 | 317 | BacillusSubtilis Pectate Lyase [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9WYR4 | 2.21e-20 | 176 | 306 | 110 | 240 | Pectate trisaccharide-lyase OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=pelA PE=1 SV=1 |
| A1CYB8 | 3.03e-20 | 164 | 395 | 96 | 320 | Probable pectate lyase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyA PE=3 SV=1 |
| B1L969 | 7.23e-20 | 176 | 306 | 108 | 238 | Pectate trisaccharide-lyase OS=Thermotoga sp. (strain RQ2) OX=126740 GN=pelA PE=3 SV=1 |
| B0XT32 | 1.39e-19 | 164 | 395 | 96 | 320 | Probable pectate lyase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyA PE=3 SV=1 |
| Q4WIT0 | 1.39e-19 | 164 | 395 | 96 | 320 | Probable pectate lyase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
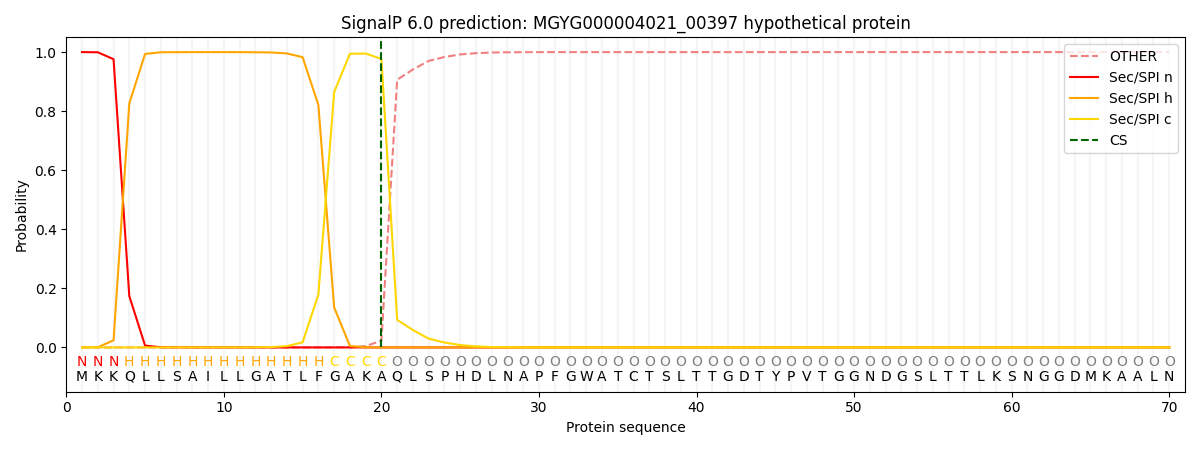
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000266 | 0.999028 | 0.000194 | 0.000173 | 0.000157 | 0.000140 |
