You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004056_01014
You are here: Home > Sequence: MGYG000004056_01014
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Alistipes_A ihumii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes_A; Alistipes_A ihumii | |||||||||||
| CAZyme ID | MGYG000004056_01014 | |||||||||||
| CAZy Family | GH127 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7442; End: 9436 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH127 | 91 | 546 | 1.8e-59 | 0.8931297709923665 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07944 | Glyco_hydro_127 | 3.58e-70 | 66 | 547 | 20 | 503 | Beta-L-arabinofuranosidase, GH127. One member of this family, from Bidobacterium longicum, UniProtKB:E8MGH8, has been characterized as an unusual beta-L-arabinofuranosidase enzyme, EC:3.2.1.185. It rleases l-arabinose from the l-arabinofuranose (Araf)-beta1,2-Araf disaccharide and also transglycosylates 1-alkanols with retention of the anomeric configuration. Terminal beta-l-arabinofuranosyl residues have been found in arabinogalactan proteins from a mumber of different plantt species. beta-l-Arabinofuranosyl linkages with 1-4 arabinofuranosides are also found in the sugar chains of extensin and solanaceous lectins, hydroxyproline (Hyp)2-rich glycoproteins that are widely observed in plant cell wall fractions. The critical residue for catalytic activity is Glu-338, in a ET/SCAS sequence context. |
| COG3533 | COG3533 | 1.71e-55 | 38 | 549 | 7 | 503 | Uncharacterized conserved protein, DUF1680 family [Function unknown]. |
| cd04792 | LanM-like | 3.18e-06 | 153 | 322 | 551 | 713 | Cyclases involved in the biosynthesis of class II lantibiotics, and similar proteins. LanM-like proteins. LanM is a bifunctional enzyme, involved in the synthesis of class II lantibiotics. It is responsible for both the dehydration and the cyclization of the precursor-peptide during lantibiotic synthesis. The C-terminal domain shows similarity to LanC, the cyclase component of the lan operon, but the N terminus seems to be unrelated to the dehydratase, LanB. |
| cd04791 | LanC_SerThrkinase | 2.57e-05 | 153 | 300 | 91 | 234 | Lanthionine synthetase C-like domain associated with serine/threonine kinases. Some members of this subgroup lack the zinc binding site and the active site residues, and therefore are most likely inactive. The function of this domain is unknown. |
| cd04434 | LanC_like | 0.003 | 88 | 321 | 38 | 270 | Cyclases involved in the biosynthesis of lantibiotics, and similar proteins. LanC is the cyclase enzyme of the lanthionine synthetase. Lanthionine is a lantibiotic, a unique class of peptide antibiotics. They are ribosomally synthesized as a precursor peptide and then post-translationally modified to contain thioether cross-links called lanthionines (Lans) or methyllanthionines (MeLans), in addition to 2,3-didehydroalanine (Dha) and (Z)-2,3-didehydrobutyrine (Dhb). These unusual amino acids are introduced by the dehydration of serine and threonine residues, followed by thioether formation via addition of cysteine thiols, catalysed by LanB and LanC or LanM. LanC, the cyclase component, is a zinc metalloprotein, whose bound metal has been proposed to activate the thiol substrate for nucleophilic addition. A related domain is also present in LanM and other pro- and eukaryotic proteins with poorly characterized functions. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ARE60508.1 | 1.83e-126 | 48 | 631 | 27 | 608 |
| SCM56172.1 | 6.46e-43 | 100 | 567 | 100 | 573 |
| QWG10444.1 | 8.28e-43 | 52 | 561 | 47 | 591 |
| QBG46791.1 | 2.83e-42 | 95 | 561 | 102 | 595 |
| AZQ63834.1 | 7.00e-42 | 89 | 561 | 91 | 591 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5MQO_A | 4.30e-26 | 52 | 560 | 49 | 620 | Glycosidehydrolase BT_1003 [Bacteroides thetaiotaomicron] |
| 3WRE_A | 2.78e-25 | 95 | 609 | 80 | 621 | Thecrystal structure of native HypBA1 from Bifidobacterium longum JCM 1217 [Bifidobacterium longum subsp. longum JCM 1217],3WRG_A The complex structure of HypBA1 with L-arabinose [Bifidobacterium longum subsp. longum JCM 1217] |
| 3WKW_A | 2.87e-25 | 95 | 609 | 80 | 621 | Crystalstructure of GH127 beta-L-arabinofuranosidase HypBA1 from Bifidobacterium longum ligand free form [Bifidobacterium longum subsp. longum JCM 1217],3WKX_A Crystal structure of GH127 beta-L-arabinofuranosidase HypBA1 from Bifidobacterium longum arabinose complex form [Bifidobacterium longum subsp. longum JCM 1217],7BZL_A Chain A, Non-reducing end beta-L-arabinofuranosidase [Bifidobacterium longum subsp. longum JCM 1217],7DIF_A Chain A, Non-reducing end beta-L-arabinofuranosidase [Bifidobacterium longum subsp. longum JCM 1217],7EXV_A Chain A, Non-reducing end beta-L-arabinofuranosidase [Bifidobacterium longum subsp. longum JCM 1217],7EXW_A Chain A, Non-reducing end beta-L-arabinofuranosidase [Bifidobacterium longum subsp. longum JCM 1217] |
| 6EX6_A | 6.30e-25 | 108 | 545 | 87 | 543 | TheGH127, Beta-arabinofuranosidase, BT3674 [Bacteroides thetaiotaomicron VPI-5482],6EX6_B The GH127, Beta-arabinofuranosidase, BT3674 [Bacteroides thetaiotaomicron VPI-5482] |
| 7EXU_A | 6.72e-25 | 95 | 609 | 80 | 621 | ChainA, Non-reducing end beta-L-arabinofuranosidase [Bifidobacterium longum subsp. longum JCM 1217] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| E8MGH8 | 1.52e-24 | 95 | 609 | 80 | 621 | Non-reducing end beta-L-arabinofuranosidase OS=Bifidobacterium longum subsp. longum (strain ATCC 15707 / DSM 20219 / JCM 1217 / NCTC 11818 / E194b) OX=565042 GN=hypBA1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
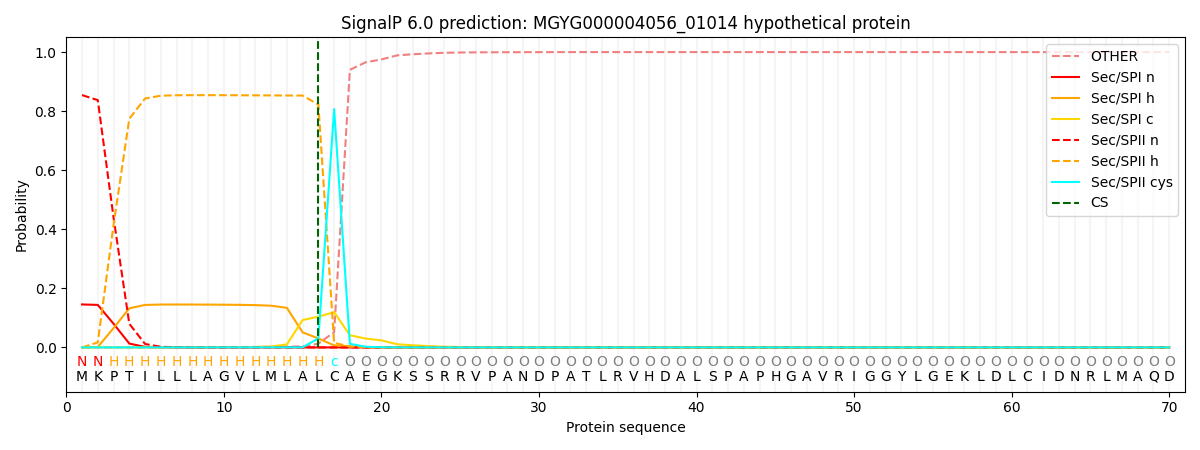
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000118 | 0.143844 | 0.855919 | 0.000047 | 0.000050 | 0.000043 |
