You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004067_00175
You are here: Home > Sequence: MGYG000004067_00175
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; CAG-274; ; | |||||||||||
| CAZyme ID | MGYG000004067_00175 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 28129; End: 30492 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 84 | 275 | 9.1e-73 | 0.9890710382513661 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam17957 | Big_7 | 9.57e-12 | 468 | 532 | 1 | 67 | Bacterial Ig domain. This entry represents a bacterial ig-like domain that is found in glycosyl hydrolase enzymes. |
| COG3866 | PelB | 7.46e-08 | 28 | 230 | 29 | 236 | Pectate lyase [Carbohydrate transport and metabolism]. |
| smart00656 | Amb_all | 1.60e-05 | 82 | 271 | 1 | 179 | Amb_all domain. |
| pfam00544 | Pec_lyase_C | 2.17e-04 | 121 | 252 | 56 | 198 | Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. |
| pfam07081 | DUF1349 | 3.03e-04 | 721 | 760 | 108 | 147 | Protein of unknown function (DUF1349). This family consists of several hypothetical bacterial proteins but contains one sequence from Saccharomyces cerevisiae. Members of this family are typically around 200 residues in length. The function of this family is unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AFH63199.1 | 3.17e-229 | 23 | 784 | 37 | 760 |
| AFC30874.1 | 3.17e-229 | 23 | 784 | 37 | 760 |
| AEI43214.1 | 6.36e-229 | 27 | 784 | 41 | 760 |
| QYR21100.1 | 4.32e-228 | 26 | 784 | 45 | 832 |
| AWV33235.1 | 5.26e-227 | 22 | 786 | 48 | 841 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3PDD_A | 3.37e-06 | 466 | 536 | 92 | 163 | ChainA, Glycoside hydrolase, family 9 [Acetivibrio thermocellus ATCC 27405] |
| 3PE9_A | 3.58e-06 | 468 | 536 | 2 | 71 | ChainA, Fibronectin(III)-like module [Acetivibrio thermocellus],3PE9_B Chain B, Fibronectin(III)-like module [Acetivibrio thermocellus],3PE9_C Chain C, Fibronectin(III)-like module [Acetivibrio thermocellus],3PE9_D Chain D, Fibronectin(III)-like module [Acetivibrio thermocellus] |
| 3PDG_A | 9.13e-06 | 468 | 534 | 2 | 69 | ChainA, Fibronectin(III)-like module [Acetivibrio thermocellus ATCC 27405] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q5B297 | 1.26e-60 | 14 | 461 | 6 | 415 | Probable pectate lyase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyC PE=3 SV=1 |
| Q0CLG7 | 1.65e-60 | 11 | 461 | 4 | 418 | Probable pectate lyase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=plyC PE=3 SV=1 |
| A1DPF0 | 3.14e-59 | 23 | 461 | 17 | 419 | Probable pectate lyase C OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyC PE=3 SV=1 |
| B0XMA2 | 5.99e-59 | 23 | 461 | 17 | 419 | Probable pectate lyase C OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyC PE=3 SV=1 |
| Q4WL88 | 8.28e-59 | 23 | 461 | 17 | 419 | Probable pectate lyase C OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
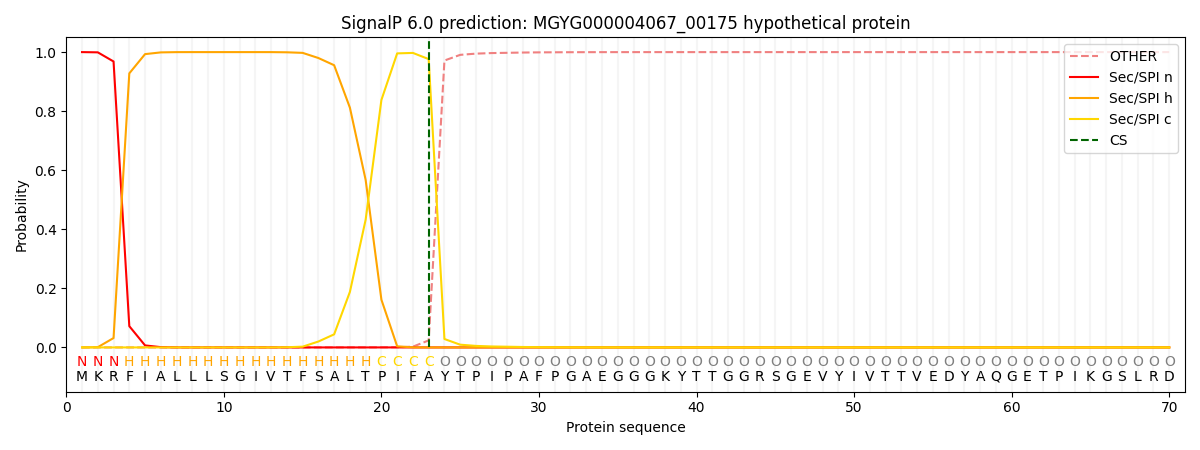
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000285 | 0.999066 | 0.000174 | 0.000167 | 0.000149 | 0.000132 |