You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004075_01794
You are here: Home > Sequence: MGYG000004075_01794
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
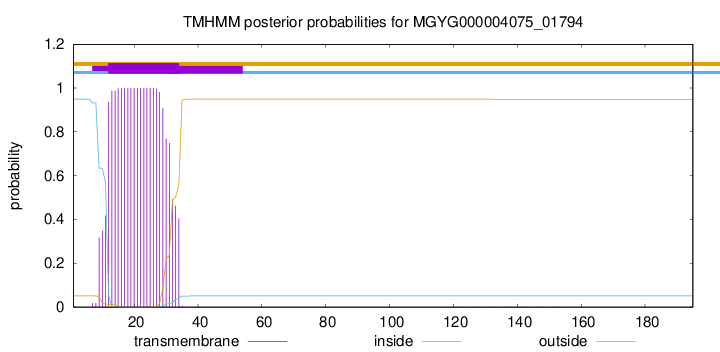
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; UMGS1591; | |||||||||||
| CAZyme ID | MGYG000004075_01794 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Soluble lytic murein transglycosylase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 25456; End: 26043 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 48 | 187 | 7.2e-30 | 0.8740740740740741 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd16896 | LT_Slt70-like | 4.87e-65 | 42 | 187 | 1 | 146 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd13401 | Slt70-like | 2.07e-46 | 40 | 191 | 1 | 152 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| cd00254 | LT-like | 2.11e-35 | 60 | 186 | 1 | 111 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd16893 | LT_MltC_MltE | 3.42e-33 | 47 | 186 | 1 | 162 | membrane-bound lytic murein transglycosylases MltC and MltE, and similar proteins. MltC and MltE are periplasmic, outer membrane attached lytic transglycosylases (LTs), which cleave beta-1,4-glycosidic bonds joining N-acetylmuramic acid and N-acetylglucosamine in the cell wall peptidoglycan, yielding 1,6-anhydromuropeptides. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda |
| pfam01464 | SLT | 4.28e-31 | 51 | 154 | 3 | 103 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCN93454.1 | 1.09e-53 | 35 | 193 | 32 | 190 |
| AYF42704.1 | 1.09e-53 | 35 | 193 | 32 | 190 |
| AYF39872.1 | 1.09e-53 | 35 | 193 | 32 | 190 |
| AVQ97208.1 | 1.09e-53 | 35 | 193 | 32 | 190 |
| ADU28212.1 | 1.09e-53 | 35 | 193 | 32 | 190 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1SLY_A | 6.92e-18 | 36 | 193 | 444 | 603 | ComplexOf The 70-Kda Soluble Lytic Transglycosylase With Bulgecin A [Escherichia coli] |
| 1QSA_A | 2.36e-17 | 36 | 193 | 444 | 603 | CrystalStructure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.65 Angstroms Resolution [Escherichia coli],1QTE_A Crystal Structure Of The 70 Kda Soluble Lytic Transglycosylase Slt70 From Escherichia Coli At 1.90 A Resolution In Complex With A 1,6- Anhydromurotripeptide [Escherichia coli] |
| 6FBT_A | 1.48e-16 | 41 | 188 | 445 | 592 | ChainA, Lytic murein transglycosylase [Pseudomonas aeruginosa] |
| 5OHU_A | 1.50e-16 | 41 | 188 | 474 | 621 | TheX-ray Structure of Lytic Transglycosylase Slt from Pseudomonas aeruginosa [Pseudomonas aeruginosa] |
| 6FCQ_A | 3.71e-16 | 41 | 188 | 445 | 592 | ChainA, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa],6FCR_A Chain A, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa],6FCS_A Chain A, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa],6FCU_A Chain A, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AGC4 | 1.31e-16 | 36 | 193 | 471 | 630 | Soluble lytic murein transglycosylase OS=Escherichia coli O157:H7 OX=83334 GN=slt PE=3 SV=1 |
| P0AGC3 | 1.31e-16 | 36 | 193 | 471 | 630 | Soluble lytic murein transglycosylase OS=Escherichia coli (strain K12) OX=83333 GN=slt PE=1 SV=1 |
| P39434 | 6.04e-16 | 36 | 193 | 471 | 630 | Soluble lytic murein transglycosylase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=slt PE=3 SV=2 |
| A7MLX9 | 1.04e-13 | 44 | 189 | 192 | 358 | Membrane-bound lytic murein transglycosylase C OS=Cronobacter sakazakii (strain ATCC BAA-894) OX=290339 GN=mltC PE=3 SV=2 |
| O31976 | 1.74e-13 | 39 | 182 | 1416 | 1538 | SPbeta prophage-derived uncharacterized transglycosylase YomI OS=Bacillus subtilis (strain 168) OX=224308 GN=yomI PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
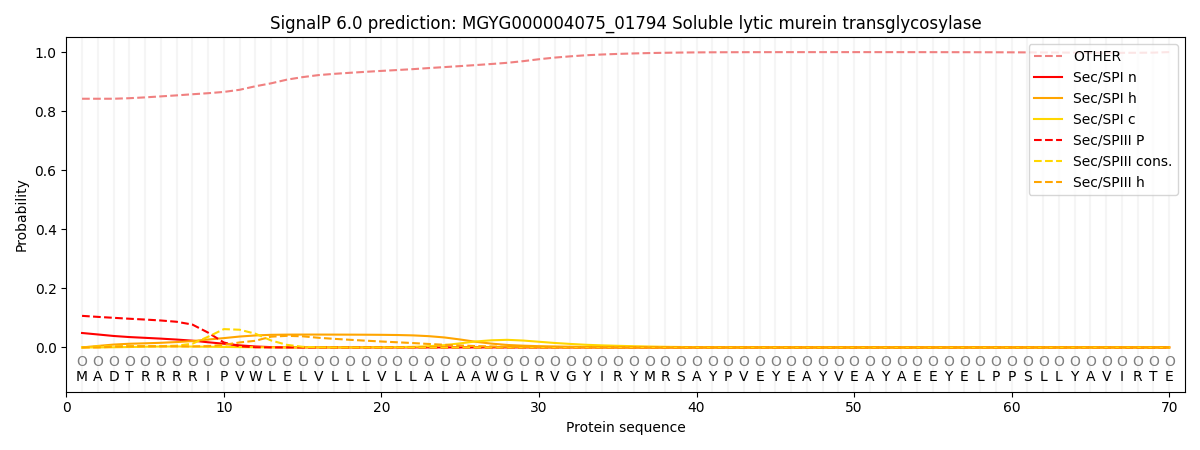
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.844101 | 0.045002 | 0.002577 | 0.000405 | 0.000242 | 0.107690 |