You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004107_00996
You are here: Home > Sequence: MGYG000004107_00996
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia_A; Christensenellales; CAG-74; Firm-11; | |||||||||||
| CAZyme ID | MGYG000004107_00996 | |||||||||||
| CAZy Family | GT39 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2339; End: 6538 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT39 | 910 | 1143 | 2.2e-53 | 0.9147982062780269 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam16192 | PMT_4TMC | 3.96e-21 | 1153 | 1317 | 1 | 170 | C-terminal four TMM region of protein-O-mannosyltransferase. PMT_4TMC is the C-terminal four membrane-pass region of protein-O-mannosyltransferases and similar enzymes. |
| COG1928 | PMT1 | 1.91e-20 | 914 | 1148 | 50 | 277 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase [Posttranslational modification, protein turnover, chaperones]. |
| pfam02366 | PMT | 1.21e-15 | 907 | 1114 | 16 | 225 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This is a family of Dolichyl-phosphate-mannose-protein mannosyltransferase proteins EC:2.4.1.109. These proteins are responsible for O-linked glycosylation of proteins, they catalyze the reaction:- Dolichyl phosphate D-mannose + protein <=> dolichyl phosphate + O-D-mannosyl-protein. Also in this family is Drosophila rotated abdomen protein which is a putative mannosyltransferase. This family appears to be distantly related to pfam02516 (A Bateman pers. obs.). This family also contains sequences from ArnTs (4-amino-4-deoxy-L-arabinose lipid A transferase). They catalyze the addition of 4-amino-4-deoxy-l-arabinose (l-Ara4N) to the lipid A moiety of the lipopolysaccharide. This is a critical modification enabling bacteria (e.g. Escherichia coli and Salmonella typhimurium) to resist killing by antimicrobial peptides such as polymyxins. Members such as undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase are predicted to have 12 trans-membrane regions. The N-terminal portion of these proteins is hypothesized to have a conserved glycosylation activity which is shared between distantly related oligosaccharyltransferases ArnT and PglB families. |
| COG4346 | COG4346 | 1.17e-14 | 930 | 1227 | 129 | 360 | Predicted membrane-bound dolichyl-phosphate-mannose-protein mannosyltransferase [Posttranslational modification, protein turnover, chaperones]. |
| COG1928 | PMT1 | 1.39e-13 | 1152 | 1295 | 498 | 637 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase [Posttranslational modification, protein turnover, chaperones]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUC68008.1 | 2.61e-309 | 2 | 1399 | 5 | 1422 |
| QTE75201.1 | 1.11e-304 | 1 | 1399 | 3 | 1422 |
| QTE71236.1 | 1.98e-304 | 16 | 1399 | 6 | 1407 |
| QUA53848.1 | 7.95e-303 | 5 | 1389 | 17 | 1362 |
| QTE68569.1 | 2.15e-302 | 2 | 1389 | 8 | 1392 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6P25_A | 1.93e-06 | 908 | 1090 | 70 | 253 | Structureof S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor and a peptide acceptor [Saccharomyces cerevisiae W303],6P2R_A Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor [Saccharomyces cerevisiae W303] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P9WN04 | 1.19e-23 | 914 | 1293 | 73 | 461 | Probable dolichyl-phosphate-mannose--protein mannosyltransferase OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=pmt PE=3 SV=2 |
| P9WN05 | 1.19e-23 | 914 | 1293 | 73 | 461 | Probable dolichyl-phosphate-mannose--protein mannosyltransferase OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=pmt PE=1 SV=2 |
| L8F4Z2 | 7.85e-20 | 903 | 1293 | 56 | 455 | Probable dolichyl-phosphate-mannose--protein mannosyltransferase OS=Mycolicibacterium smegmatis (strain MKD8) OX=1214915 GN=pmt PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
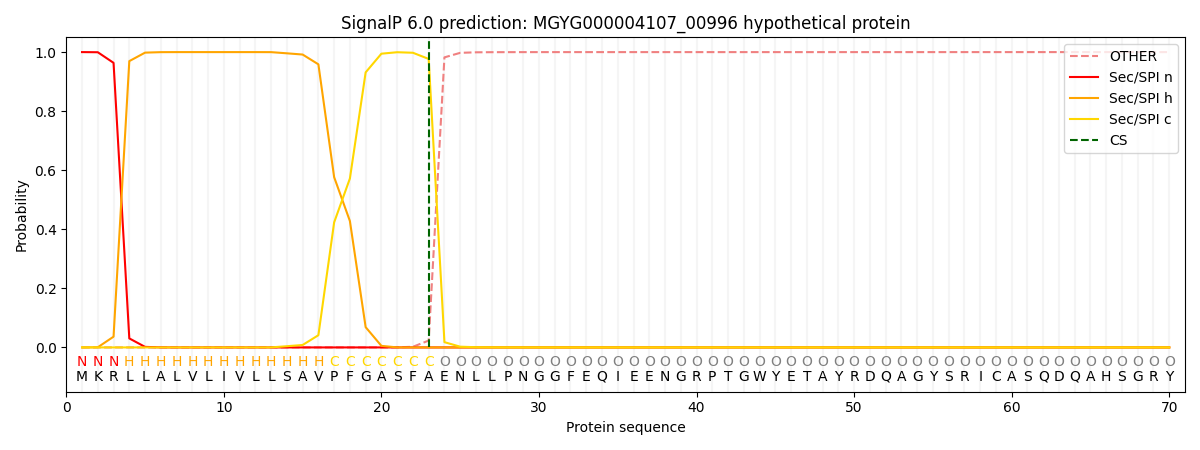
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000205 | 0.999175 | 0.000164 | 0.000166 | 0.000146 | 0.000136 |
TMHMM Annotations download full data without filtering help
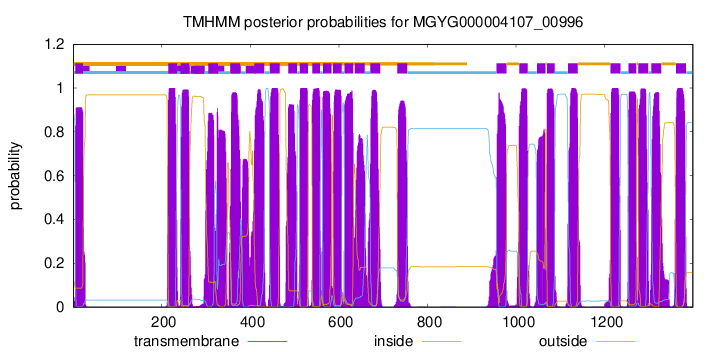
| start | end |
|---|---|
| 5 | 22 |
| 215 | 234 |
| 243 | 262 |
| 305 | 327 |
| 355 | 377 |
| 409 | 431 |
| 444 | 466 |
| 486 | 505 |
| 512 | 530 |
| 540 | 557 |
| 564 | 583 |
| 587 | 606 |
| 613 | 632 |
| 636 | 658 |
| 671 | 693 |
| 732 | 754 |
| 956 | 978 |
| 1007 | 1026 |
| 1047 | 1066 |
| 1070 | 1087 |
| 1117 | 1139 |
| 1213 | 1235 |
| 1254 | 1271 |
| 1276 | 1298 |
| 1305 | 1327 |
| 1361 | 1383 |
