You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004188_01504
You are here: Home > Sequence: MGYG000004188_01504
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; | |||||||||||
| CAZyme ID | MGYG000004188_01504 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 39458; End: 41107 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 68 | 253 | 3.5e-50 | 0.5535714285714286 |
| GH5 | 212 | 449 | 1.6e-17 | 0.5874125874125874 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 2.28e-21 | 68 | 450 | 2 | 270 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 4.81e-07 | 76 | 242 | 55 | 191 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRO24956.1 | 1.24e-201 | 39 | 549 | 13 | 522 |
| QDM45840.1 | 2.43e-118 | 54 | 539 | 9 | 493 |
| QTH41332.1 | 1.62e-115 | 59 | 545 | 4 | 492 |
| QJD87094.1 | 1.43e-113 | 59 | 545 | 4 | 492 |
| QNK55040.1 | 7.79e-113 | 59 | 545 | 4 | 491 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2OSW_A | 1.87e-36 | 51 | 505 | 17 | 438 | Endo-glycoceramidaseII from Rhodococcus sp. [Rhodococcus sp.],2OSW_B Endo-glycoceramidase II from Rhodococcus sp. [Rhodococcus sp.],2OYK_A Endo-glycoceramidase II from Rhodococcus sp.: cellobiose-like isofagomine complex [Rhodococcus sp.],2OYK_B Endo-glycoceramidase II from Rhodococcus sp.: cellobiose-like isofagomine complex [Rhodococcus sp.],2OYL_A Endo-glycoceramidase II from Rhodococcus sp.: cellobiose-like imidazole complex [Rhodococcus sp.],2OYL_B Endo-glycoceramidase II from Rhodococcus sp.: cellobiose-like imidazole complex [Rhodococcus sp.],2OYM_A Endo-glycoceramidase II from Rhodococcus sp.: five-membered iminocyclitol complex [Rhodococcus sp.],2OYM_B Endo-glycoceramidase II from Rhodococcus sp.: five-membered iminocyclitol complex [Rhodococcus sp.] |
| 2OSX_A | 8.92e-36 | 51 | 505 | 17 | 438 | ChainA, Endoglycoceramidase II [Rhodococcus sp.] |
| 2OSY_A | 1.22e-35 | 51 | 505 | 17 | 438 | ChainA, Endoglycoceramidase II [Rhodococcus sp.],2OSY_B Chain B, Endoglycoceramidase II [Rhodococcus sp.] |
| 5CCU_A | 1.73e-23 | 58 | 511 | 43 | 441 | ChainA, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5CCU_B Chain B, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S] |
| 5J14_A | 7.45e-23 | 58 | 511 | 43 | 441 | ChainA, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5J14_B Chain B, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5J7Z_A Chain A, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S],5J7Z_B Chain B, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A0A3S5YBC7 | 8.99e-23 | 58 | 511 | 35 | 433 | Endoglycoceramidase I OS=Rhodococcus hoagii (strain 103S) OX=685727 GN=REQ_38260 PE=1 SV=1 |
| Q9GV16 | 7.55e-18 | 65 | 245 | 40 | 243 | Endoglycoceramidase OS=Cyanea nozakii OX=135523 PE=1 SV=1 |
| Q6L6S1 | 5.71e-16 | 66 | 263 | 32 | 251 | Endoglycoceramidase OS=Hydra vulgaris OX=6087 PE=1 SV=1 |
| I1BTD7 | 3.80e-13 | 63 | 245 | 19 | 258 | Glucosylceramidase OS=Rhizopus delemar (strain RA 99-880 / ATCC MYA-4621 / FGSC 9543 / NRRL 43880) OX=246409 GN=ERC1 PE=1 SV=1 |
| H1AE13 | 7.13e-13 | 59 | 258 | 6 | 254 | Glucosylceramidase OS=Neosartorya fumigata OX=746128 GN=egc1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
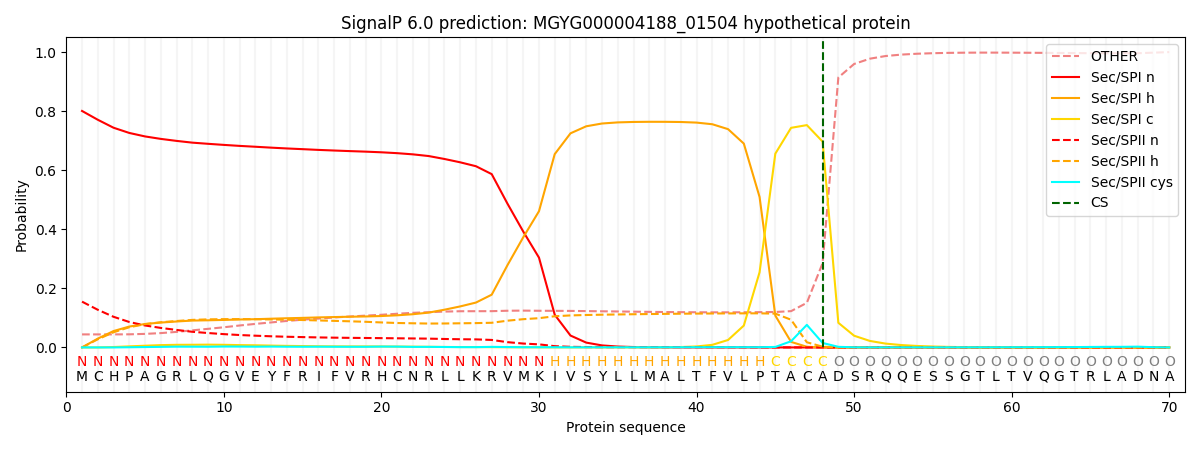
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.048176 | 0.771750 | 0.178418 | 0.000572 | 0.000557 | 0.000513 |
