You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004208_02214
You are here: Home > Sequence: MGYG000004208_02214
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-485 sp900759795 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; CAG-485; CAG-485 sp900759795 | |||||||||||
| CAZyme ID | MGYG000004208_02214 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | Xyloglucan-specific endo-beta-1,4-glucanase BoGH5A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 4324; End: 5820 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 169 | 462 | 7.8e-100 | 0.9927536231884058 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 1.03e-54 | 163 | 461 | 10 | 269 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 4.95e-22 | 147 | 466 | 45 | 365 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| cd14948 | BACON | 8.02e-16 | 25 | 111 | 1 | 83 | Bacteroidetes-Associated Carbohydrate-binding (putative) Often N-terminal (BACON) domain. The BACON domain is found in diverse domain architectures and accociated with a wide variety of domains, including carbohydrate-active enzymes and proteases. It was named for its suggested function of carbohydrate binding; the latter was inferred from domain architectures, sequence conservation, and phyletic distribution. However, recent experimental data suggest that its primary function in Bacteroides ovatus endo-xyloglucanase BoGH5A is to distance the catalytic module from the cell surface and confer additional mobility to the catalytic domain for attack of the polysaccharide. No evidence for a direct role in carbohydrate binding could be found in that case. The large majority of BACON domains are found in Bacteroidetes. |
| pfam13004 | BACON | 1.59e-08 | 53 | 111 | 4 | 61 | Putative binding domain, N-terminal. The BACON (Bacteroidetes-Associated Carbohydrate-binding Often N-terminal) domain is an all-beta domain found in diverse architectures, principally in combination with carbohydrate-active enzymes and proteases. These architectures suggest a carbohydrate-binding function which is also supported by the nature of BACON's few conserved amino-acids. The phyletic distribution of BACON and other data tentatively suggest that it may frequently function to bind mucin. Further work with the characterized structure of a member of glycoside hydrolase family 5 enzyme, Structure 3ZMR, has found no evidence for carbohydrate-binding for this domain. |
| pfam19190 | BACON_2 | 2.67e-06 | 53 | 113 | 33 | 91 | Viral BACON domain. This family represents a distinct class of BACON domains found in crAss-like phages, the most common viral family in the human gut, in which they are found in tail fiber genes. This suggests they may play a role in phage-host interactions. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIK55671.1 | 2.54e-107 | 2 | 488 | 6 | 486 |
| QIK61079.1 | 7.16e-107 | 2 | 488 | 6 | 486 |
| ASB50110.1 | 1.44e-103 | 17 | 491 | 103 | 573 |
| QRK07516.1 | 1.44e-101 | 146 | 494 | 34 | 374 |
| QEC45584.1 | 1.53e-101 | 16 | 477 | 15 | 460 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OYC_A | 4.96e-78 | 135 | 491 | 34 | 393 | GH5endo-xyloglucanase from Cellvibrio japonicus [Cellvibrio japonicus Ueda107],5OYC_B GH5 endo-xyloglucanase from Cellvibrio japonicus [Cellvibrio japonicus Ueda107],5OYD_A GH5 endo-xyloglucanase from Cellvibrio japonicus [Cellvibrio japonicus Ueda107],5OYD_B GH5 endo-xyloglucanase from Cellvibrio japonicus [Cellvibrio japonicus Ueda107],5OYE_A GH5 endo-xyloglucanase from Cellvibrio japonicus [Cellvibrio japonicus Ueda107],5OYE_B GH5 endo-xyloglucanase from Cellvibrio japonicus [Cellvibrio japonicus Ueda107] |
| 6HA9_A | 3.86e-77 | 135 | 491 | 34 | 393 | Structureof an endo-Xyloglucanase from Cellvibrio japonicus complexed with XXXG(2F)-beta-DNP [Cellvibrio japonicus Ueda107],6HA9_B Structure of an endo-Xyloglucanase from Cellvibrio japonicus complexed with XXXG(2F)-beta-DNP [Cellvibrio japonicus Ueda107],6HAA_A Structure of a covalent complex of endo-Xyloglucanase from Cellvibrio japonicus after reacting with XXXG(2F)-beta-DNP [Cellvibrio japonicus Ueda107],6HAA_B Structure of a covalent complex of endo-Xyloglucanase from Cellvibrio japonicus after reacting with XXXG(2F)-beta-DNP [Cellvibrio japonicus Ueda107] |
| 2JEP_A | 6.83e-74 | 146 | 490 | 35 | 393 | Nativefamily 5 xyloglucanase from Paenibacillus pabuli [Paenibacillus pabuli],2JEP_B Native family 5 xyloglucanase from Paenibacillus pabuli [Paenibacillus pabuli],2JEQ_A Family 5 xyloglucanase from Paenibacillus pabuli in complex with ligand [Paenibacillus pabuli] |
| 3ZMR_A | 7.68e-74 | 21 | 490 | 3 | 465 | Bacteroidesovatus GH5 xyloglucanase in complex with a XXXG heptasaccharide [Bacteroides ovatus],3ZMR_B Bacteroides ovatus GH5 xyloglucanase in complex with a XXXG heptasaccharide [Bacteroides ovatus] |
| 6WQY_A | 2.19e-72 | 143 | 478 | 21 | 367 | ChainA, Cellulase [Phocaeicola salanitronis DSM 18170] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| A7LXT7 | 4.82e-76 | 1 | 490 | 20 | 500 | Xyloglucan-specific endo-beta-1,4-glucanase BoGH5A OS=Bacteroides ovatus (strain ATCC 8483 / DSM 1896 / JCM 5824 / BCRC 10623 / CCUG 4943 / NCTC 11153) OX=411476 GN=BACOVA_02653 PE=1 SV=1 |
| O08342 | 1.40e-70 | 145 | 490 | 39 | 398 | Endoglucanase A OS=Paenibacillus barcinonensis OX=198119 GN=celA PE=1 SV=1 |
| P28621 | 4.34e-64 | 145 | 491 | 41 | 374 | Endoglucanase B OS=Clostridium cellulovorans (strain ATCC 35296 / DSM 3052 / OCM 3 / 743B) OX=573061 GN=engB PE=3 SV=1 |
| P28623 | 3.60e-63 | 146 | 490 | 43 | 371 | Endoglucanase D OS=Clostridium cellulovorans (strain ATCC 35296 / DSM 3052 / OCM 3 / 743B) OX=573061 GN=engD PE=1 SV=2 |
| P23660 | 3.13e-59 | 145 | 475 | 26 | 347 | Endoglucanase A OS=Ruminococcus albus OX=1264 GN=celA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
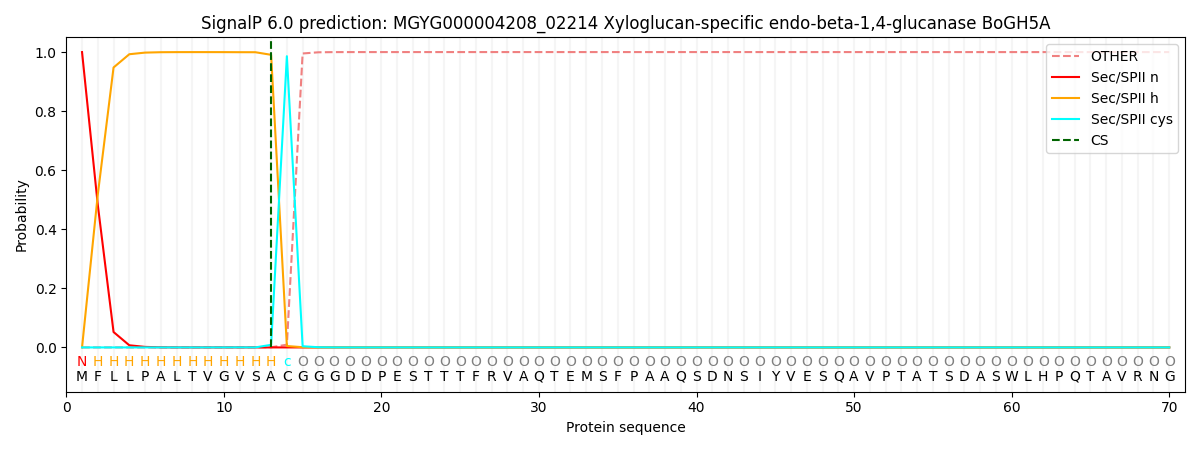
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000002 | 1.000063 | 0.000000 | 0.000000 | 0.000000 |
