You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004248_00916
You are here: Home > Sequence: MGYG000004248_00916
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-170 sp000436735 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Oscillospiraceae; CAG-170; CAG-170 sp000436735 | |||||||||||
| CAZyme ID | MGYG000004248_00916 | |||||||||||
| CAZy Family | GH123 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 61873; End: 67587 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH123 | 50 | 580 | 3.9e-43 | 0.9237918215613383 |
| CBM66 | 1223 | 1363 | 1.1e-16 | 0.8580645161290322 |
| CBM66 | 1028 | 1169 | 5.4e-16 | 0.896774193548387 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam09479 | Flg_new | 4.04e-16 | 1645 | 1706 | 2 | 65 | Listeria-Bacteroides repeat domain (List_Bact_rpt). This model describes a conserved core region of about 43 residues, which occurs in at least two families of tandem repeats. These include 78-residue repeats which occur from 2 to 15 times in some proteins of Bacteroides forsythus ATCC 43037, and 70-residue repeats found in families of internalins of Listeria species. Single copies are found in proteins of Fibrobacter succinogenes, Geobacter sulfurreducens, and a few other bacteria. |
| NF033190 | inl_like_NEAT_1 | 5.82e-11 | 1730 | 1897 | 582 | 748 | NEAT domain-containing leucine-rich repeat protein. Members of this family have an N-terminal NEAT (near transporter) domain often associated with iron transport, followed by a leucine-rich repeat region with significant sequence similarity to the internalins of Listeria monocytogenes. However, since Bacillus cereus (from which this protein was described, in PMID:16978259) is not considered an intracellular pathogen, and the function may be iron transport rather than internalization, applying the name "internalin" to this family probably would be misleading. |
| pfam00395 | SLH | 2.14e-08 | 1730 | 1769 | 1 | 40 | S-layer homology domain. |
| pfam13320 | DUF4091 | 3.10e-07 | 502 | 562 | 1 | 65 | Domain of unknown function (DUF4091). This presumed domain is functionally uncharacterized. This domain family is found in bacteria, archaea and eukaryotes, and is approximately 70 amino acids in length. There is a single completely conserved residue G that may be functionally important. |
| pfam00395 | SLH | 6.66e-07 | 1790 | 1830 | 1 | 41 | S-layer homology domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QNE29218.1 | 7.14e-153 | 13 | 812 | 19 | 786 |
| QEV44207.1 | 3.23e-144 | 33 | 813 | 9 | 766 |
| QJD85658.1 | 8.71e-142 | 22 | 981 | 27 | 934 |
| QTH41935.1 | 8.20e-66 | 1 | 624 | 1 | 753 |
| BBI36344.1 | 2.66e-58 | 43 | 621 | 44 | 768 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P19424 | 2.34e-14 | 1717 | 1901 | 28 | 214 | Endoglucanase OS=Bacillus sp. (strain KSM-635) OX=1415 PE=1 SV=1 |
| P38535 | 1.68e-13 | 1723 | 1903 | 897 | 1083 | Exoglucanase XynX OS=Acetivibrio thermocellus OX=1515 GN=xynX PE=3 SV=1 |
| P38536 | 1.73e-12 | 1730 | 1903 | 1682 | 1857 | Amylopullulanase OS=Thermoanaerobacterium thermosulfurigenes OX=33950 GN=amyB PE=3 SV=2 |
| P38537 | 1.32e-11 | 1736 | 1904 | 40 | 209 | Surface-layer 125 kDa protein OS=Lysinibacillus sphaericus OX=1421 PE=3 SV=1 |
| P36917 | 9.80e-08 | 1723 | 1831 | 1045 | 1155 | Endo-1,4-beta-xylanase A OS=Thermoanaerobacterium saccharolyticum OX=28896 GN=xynA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
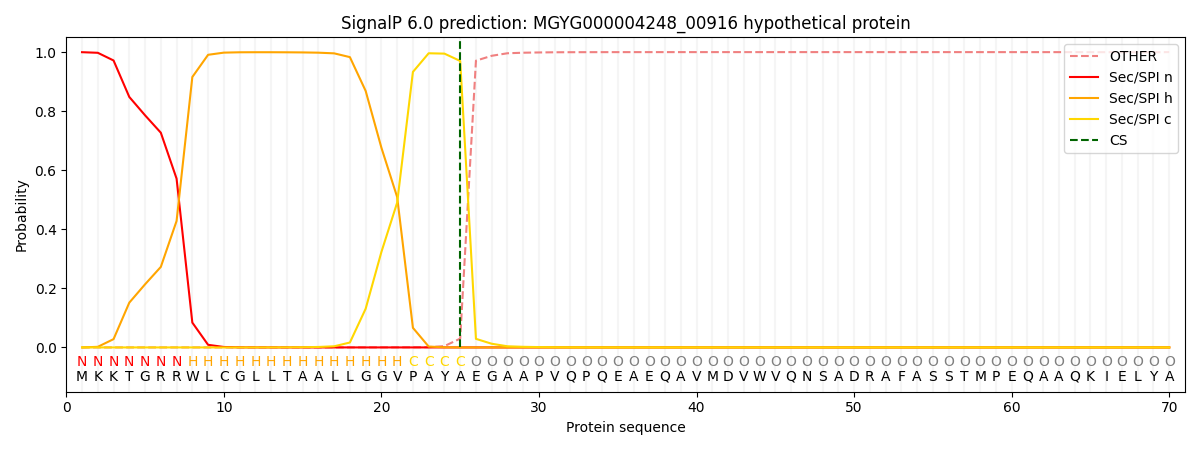
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000810 | 0.997542 | 0.000394 | 0.000724 | 0.000303 | 0.000205 |
