You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004275_00955
You are here: Home > Sequence: MGYG000004275_00955
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruminococcus_C sp000437175 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminococcus_C; Ruminococcus_C sp000437175 | |||||||||||
| CAZyme ID | MGYG000004275_00955 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 7795; End: 10926 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 212 | 436 | 5.8e-32 | 0.9356435643564357 |
| CBM13 | 558 | 704 | 3.5e-19 | 0.7021276595744681 |
| CBM13 | 712 | 877 | 1.7e-18 | 0.8191489361702128 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3866 | PelB | 2.73e-31 | 160 | 510 | 17 | 341 | Pectate lyase [Carbohydrate transport and metabolism]. |
| smart00656 | Amb_all | 5.01e-18 | 239 | 437 | 17 | 189 | Amb_all domain. |
| pfam14200 | RicinB_lectin_2 | 4.09e-16 | 643 | 742 | 1 | 89 | Ricin-type beta-trefoil lectin domain-like. |
| pfam14200 | RicinB_lectin_2 | 4.92e-16 | 556 | 637 | 12 | 89 | Ricin-type beta-trefoil lectin domain-like. |
| pfam14200 | RicinB_lectin_2 | 2.57e-14 | 597 | 686 | 5 | 89 | Ricin-type beta-trefoil lectin domain-like. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL17651.1 | 9.04e-160 | 1 | 699 | 1 | 687 |
| CDM70399.1 | 2.52e-141 | 30 | 703 | 24 | 697 |
| AUO18238.1 | 2.30e-137 | 33 | 686 | 31 | 692 |
| CBL16867.1 | 1.46e-111 | 29 | 854 | 30 | 851 |
| ASR46637.1 | 4.42e-68 | 19 | 525 | 20 | 536 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3ZSC_A | 6.99e-15 | 190 | 406 | 20 | 214 | Catalyticfunction and substrate recognition of the pectate lyase from Thermotoga maritima [Thermotoga maritima] |
| 1VBL_A | 2.04e-10 | 291 | 405 | 192 | 301 | Structureof the thermostable pectate lyase PL 47 [Bacillus sp. TS-47] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P94449 | 3.77e-24 | 173 | 516 | 35 | 342 | Pectin lyase OS=Bacillus subtilis OX=1423 GN=pelB PE=1 SV=1 |
| O34819 | 6.86e-24 | 173 | 516 | 35 | 342 | Pectin lyase OS=Bacillus subtilis (strain 168) OX=224308 GN=pelB PE=3 SV=1 |
| P27027 | 1.79e-16 | 171 | 516 | 8 | 309 | Pectin lyase OS=Pseudomonas marginalis OX=298 GN=pnl PE=1 SV=2 |
| Q9WYR4 | 2.05e-15 | 190 | 406 | 47 | 241 | Pectate trisaccharide-lyase OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=pelA PE=1 SV=1 |
| B1L969 | 6.45e-15 | 190 | 406 | 45 | 239 | Pectate trisaccharide-lyase OS=Thermotoga sp. (strain RQ2) OX=126740 GN=pelA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
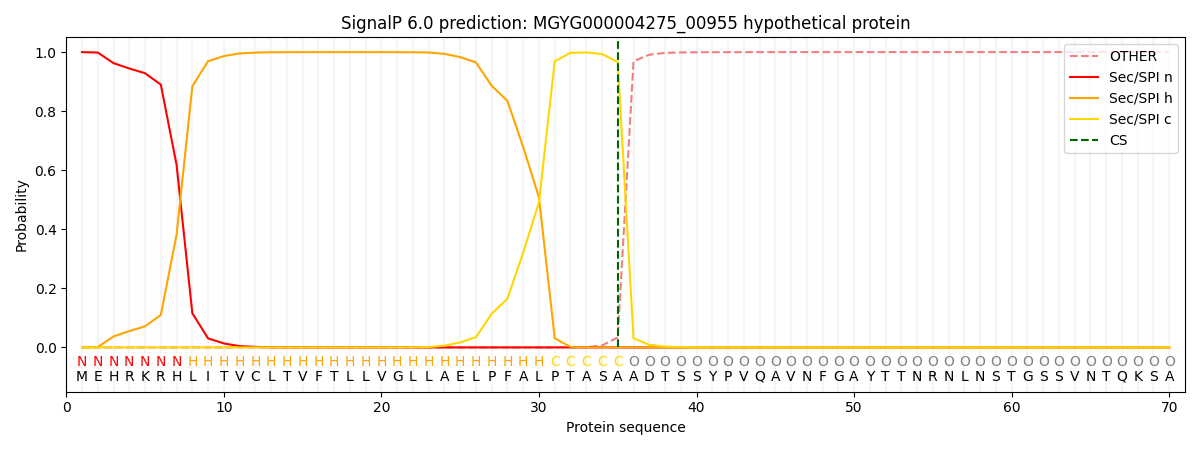
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000398 | 0.998846 | 0.000181 | 0.000221 | 0.000168 | 0.000150 |