You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004329_00715
You are here: Home > Sequence: MGYG000004329_00715
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotellamassilia sp900539625 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotellamassilia; Prevotellamassilia sp900539625 | |||||||||||
| CAZyme ID | MGYG000004329_00715 | |||||||||||
| CAZy Family | GH88 | |||||||||||
| CAZyme Description | Unsaturated chondroitin disaccharide hydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 38497; End: 39729 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH88 | 71 | 397 | 4.4e-122 | 0.9756838905775076 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07470 | Glyco_hydro_88 | 1.28e-08 | 72 | 270 | 28 | 207 | Glycosyl Hydrolase Family 88. Unsaturated glucuronyl hydrolase catalyzes the hydrolytic release of unsaturated glucuronic acids from oligosaccharides (EC:3.2.1.-) produced by the reactions of polysaccharide lyases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALO49667.1 | 7.25e-186 | 1 | 407 | 1 | 405 |
| QCT80147.1 | 1.73e-184 | 27 | 402 | 15 | 390 |
| BAD47132.1 | 1.73e-184 | 27 | 402 | 15 | 390 |
| CUA16984.1 | 2.65e-184 | 27 | 402 | 27 | 402 |
| ANQ62645.1 | 2.65e-184 | 27 | 402 | 27 | 402 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3WIW_A | 6.69e-97 | 44 | 401 | 44 | 391 | Crystalstructure of unsaturated glucuronyl hydrolase specific for heparin [Pedobacter heparinus DSM 2366] |
| 1VD5_A | 8.49e-42 | 72 | 397 | 41 | 363 | CrystalStructure of Unsaturated Glucuronyl Hydrolase, Responsible for the Degradation of Glycosaminoglycan, from Bacillus sp. GL1 at 1.8 A Resolution [Bacillus sp. GL1],2D5J_A Unsaturated Glucuronyl Hydrolase Triggers Hydration of Vinyl Ether Group but not of Glycosidic Bond [Bacillus sp. GL1],2D5J_B Unsaturated Glucuronyl Hydrolase Triggers Hydration of Vinyl Ether Group but not of Glycosidic Bond [Bacillus sp. GL1],2FUZ_A UGL hexagonal crystal structure without glycine and DTT molecules [Bacillus sp. GL1] |
| 2AHF_A | 4.45e-41 | 72 | 397 | 41 | 363 | ChainA, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2AHF_B Chain B, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2AHG_A Chain A, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2AHG_B Chain B, unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV0_A Chain A, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV0_B Chain B, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV1_A Chain A, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1],2FV1_B Chain B, Unsaturated glucuronyl hydrolase [Bacillus sp. GL1] |
| 2ZZR_A | 6.70e-39 | 46 | 397 | 31 | 388 | Crystalstructure of unsaturated glucuronyl hydrolase from Streptcoccus agalactiae [Streptococcus agalactiae] |
| 3ANJ_A | 6.83e-39 | 46 | 397 | 32 | 389 | Crystalstructure of unsaturated glucuronyl hydrolase from Streptcoccus agalactiae [Streptococcus agalactiae serogroup III] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| T2KLZ3 | 3.65e-104 | 72 | 403 | 76 | 399 | Unsaturated glucuronyl hydrolase OS=Formosa agariphila (strain DSM 15362 / KCTC 12365 / LMG 23005 / KMM 3901 / M-2Alg 35-1) OX=1347342 GN=BN863_21900 PE=1 SV=1 |
| Q9A0T3 | 3.57e-44 | 29 | 397 | 30 | 390 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus pyogenes serotype M1 OX=301447 GN=ugl PE=1 SV=1 |
| Q9RC92 | 4.65e-41 | 72 | 397 | 41 | 363 | Unsaturated glucuronyl hydrolase OS=Bacillus sp. (strain GL1) OX=84635 GN=ugl PE=1 SV=1 |
| Q8E372 | 3.74e-38 | 46 | 397 | 32 | 389 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus agalactiae serotype III (strain NEM316) OX=211110 GN=gbs1889 PE=1 SV=1 |
| Q8DR77 | 6.94e-38 | 72 | 397 | 66 | 387 | Unsaturated chondroitin disaccharide hydrolase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=ugl PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
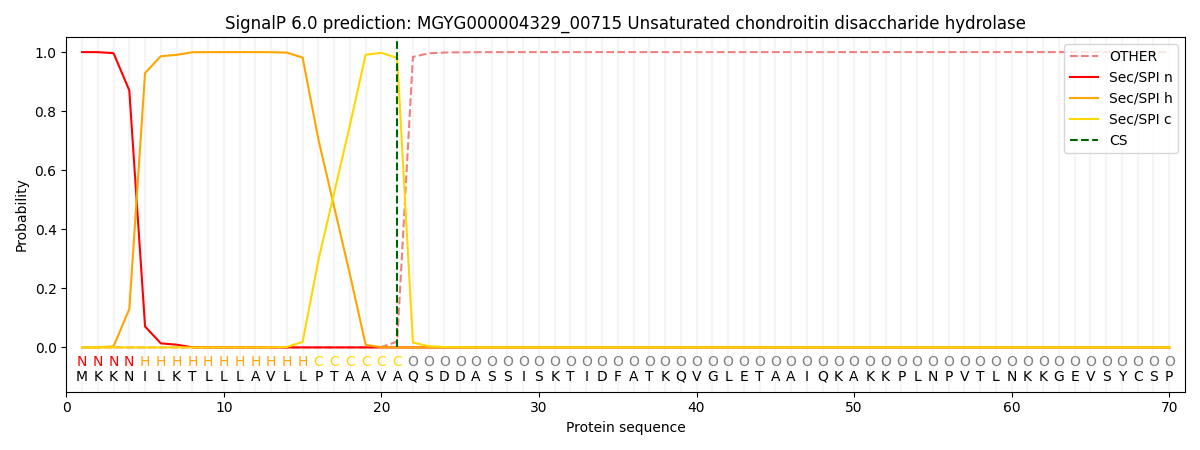
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000284 | 0.998974 | 0.000166 | 0.000188 | 0.000166 | 0.000161 |
