You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004479_01043
You are here: Home > Sequence: MGYG000004479_01043
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola mediterraneensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola mediterraneensis | |||||||||||
| CAZyme ID | MGYG000004479_01043 | |||||||||||
| CAZy Family | GH105 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 21827; End: 23896 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd01827 | sialate_O-acetylesterase_like1 | 2.74e-85 | 24 | 215 | 1 | 188 | sialate O-acetylesterase_like family of the SGNH hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| pfam13472 | Lipase_GDSL_2 | 6.49e-28 | 28 | 205 | 1 | 176 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
| pfam03629 | SASA | 5.79e-23 | 302 | 582 | 2 | 226 | Carbohydrate esterase, sialic acid-specific acetylesterase. The catalytic triad of this esterase enzyme comprises residues Ser127, His403 and Asp391 in UniProtKB:P70665. |
| cd00229 | SGNH_hydrolase | 9.99e-23 | 26 | 213 | 1 | 187 | SGNH_hydrolase, or GDSL_hydrolase, is a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the typical Ser-His-Asp(Glu) triad from other serine hydrolases, but may lack the carboxlic acid. |
| cd01834 | SGNH_hydrolase_like_2 | 6.10e-18 | 23 | 209 | 1 | 187 | SGNH_hydrolase subfamily. SGNH hydrolases are a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB42485.1 | 2.67e-239 | 22 | 684 | 29 | 692 |
| AHW60353.1 | 4.15e-218 | 19 | 684 | 31 | 700 |
| AWI09525.1 | 2.58e-90 | 221 | 685 | 636 | 1123 |
| QDT62262.1 | 3.37e-89 | 192 | 684 | 917 | 1438 |
| QUT73831.1 | 3.54e-75 | 219 | 685 | 22 | 471 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7KMM_A | 1.45e-51 | 223 | 685 | 25 | 632 | ChainA, Sialic acid-specific 9-O-acetylesterase [Xanthomonas citri pv. citri str. 306],7KMM_B Chain B, Sialic acid-specific 9-O-acetylesterase [Xanthomonas citri pv. citri str. 306] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P82450 | 2.76e-34 | 220 | 630 | 25 | 469 | Sialate O-acetylesterase OS=Rattus norvegicus OX=10116 GN=Siae PE=1 SV=2 |
| P70665 | 3.21e-32 | 220 | 598 | 25 | 437 | Sialate O-acetylesterase OS=Mus musculus OX=10090 GN=Siae PE=1 SV=3 |
| Q5RFU0 | 1.39e-28 | 220 | 612 | 25 | 425 | Sialate O-acetylesterase OS=Pongo abelii OX=9601 GN=SIAE PE=2 SV=1 |
| Q9HAT2 | 1.86e-28 | 220 | 612 | 25 | 425 | Sialate O-acetylesterase OS=Homo sapiens OX=9606 GN=SIAE PE=1 SV=1 |
| D5EV35 | 7.99e-25 | 17 | 214 | 267 | 479 | Acetylxylan esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axeA1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
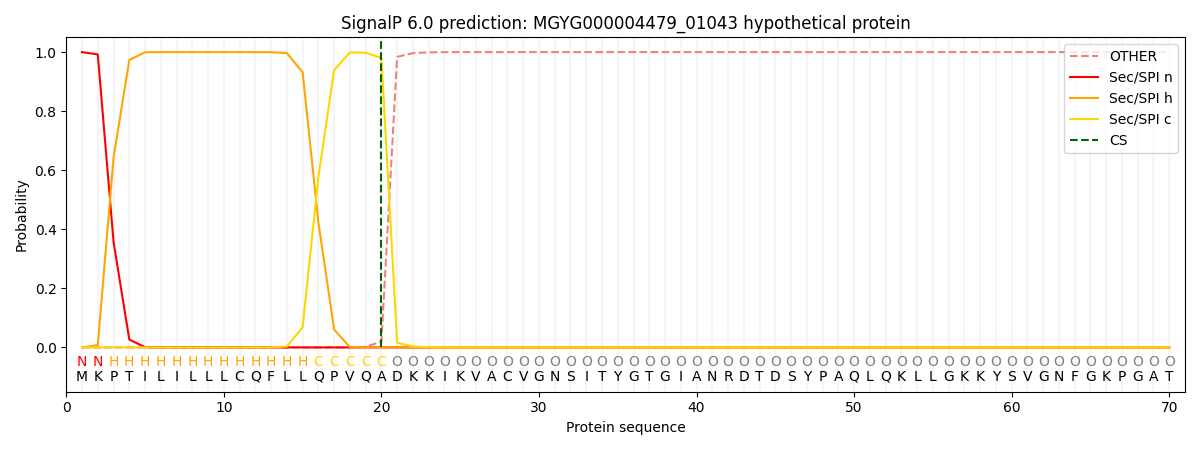
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000200 | 0.999217 | 0.000167 | 0.000133 | 0.000133 | 0.000128 |
