You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004499_03045
You are here: Home > Sequence: MGYG000004499_03045
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Victivallis sp900551245 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; Victivallaceae; Victivallis; Victivallis sp900551245 | |||||||||||
| CAZyme ID | MGYG000004499_03045 | |||||||||||
| CAZy Family | GH63 | |||||||||||
| CAZyme Description | Cytoplasmic trehalase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 12955; End: 14433 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10137 | PRK10137 | 8.68e-16 | 234 | 361 | 607 | 742 | alpha-glucosidase; Provisional |
| COG1626 | TreA | 4.27e-13 | 225 | 374 | 370 | 507 | Neutral trehalase [Carbohydrate transport and metabolism]. |
| pfam01204 | Trehalase | 6.59e-12 | 228 | 383 | 324 | 478 | Trehalase. Trehalase (EC:3.2.1.28) is known to recycle trehalose to glucose. Trehalose is a physiological hallmark of heat-shock response in yeast and protects of proteins and membranes against a variety of stresses. This family is found in conjunction with pfam07492 in fungi. |
| pfam03200 | Glyco_hydro_63 | 7.67e-10 | 186 | 407 | 235 | 487 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| PRK13270 | treF | 7.65e-09 | 232 | 378 | 376 | 501 | alpha,alpha-trehalase TreF. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QZT38129.1 | 2.30e-146 | 2 | 482 | 22 | 496 |
| AHF90735.1 | 3.22e-143 | 18 | 487 | 27 | 510 |
| AVM44762.1 | 2.27e-140 | 18 | 486 | 98 | 593 |
| QWO96172.1 | 2.16e-139 | 27 | 482 | 52 | 506 |
| QWP70936.1 | 2.16e-139 | 27 | 482 | 52 | 506 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4WVA_A | 1.99e-09 | 250 | 354 | 277 | 378 | Crystalstructure of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8 in complex with Tris [Thermus thermophilus HB8],4WVA_B Crystal structure of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8 in complex with Tris [Thermus thermophilus HB8],4WVB_A Crystal structure of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8 in complex with glucose [Thermus thermophilus HB8],4WVB_B Crystal structure of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8 in complex with glucose [Thermus thermophilus HB8],4WVC_A Crystal structure of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8 in complex with Tris and D-glycerate [Thermus thermophilus HB8],4WVC_B Crystal structure of GH63 mannosylglycerate hydrolase from Thermus thermophilus HB8 in complex with Tris and D-glycerate [Thermus thermophilus HB8] |
| 2Z07_A | 1.08e-08 | 250 | 354 | 277 | 378 | Crystalstructure of uncharacterized conserved protein from Thermus thermophilus HB8 [Thermus thermophilus],2Z07_B Crystal structure of uncharacterized conserved protein from Thermus thermophilus HB8 [Thermus thermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O42622 | 4.60e-07 | 245 | 382 | 526 | 663 | Cytosolic neutral trehalase OS=Magnaporthe oryzae (strain 70-15 / ATCC MYA-4617 / FGSC 8958) OX=242507 GN=NTH1 PE=2 SV=2 |
| Q757L1 | 1.06e-06 | 225 | 382 | 505 | 666 | Probable trehalase OS=Ashbya gossypii (strain ATCC 10895 / CBS 109.51 / FGSC 9923 / NRRL Y-1056) OX=284811 GN=NTH2 PE=3 SV=1 |
| C0Q162 | 1.21e-06 | 231 | 360 | 375 | 495 | Cytoplasmic trehalase OS=Salmonella paratyphi C (strain RKS4594) OX=476213 GN=treF PE=3 SV=1 |
| B5RGS3 | 1.21e-06 | 231 | 360 | 375 | 495 | Cytoplasmic trehalase OS=Salmonella gallinarum (strain 287/91 / NCTC 13346) OX=550538 GN=treF PE=3 SV=1 |
| Q57IL9 | 1.21e-06 | 231 | 360 | 375 | 495 | Cytoplasmic trehalase OS=Salmonella choleraesuis (strain SC-B67) OX=321314 GN=treF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
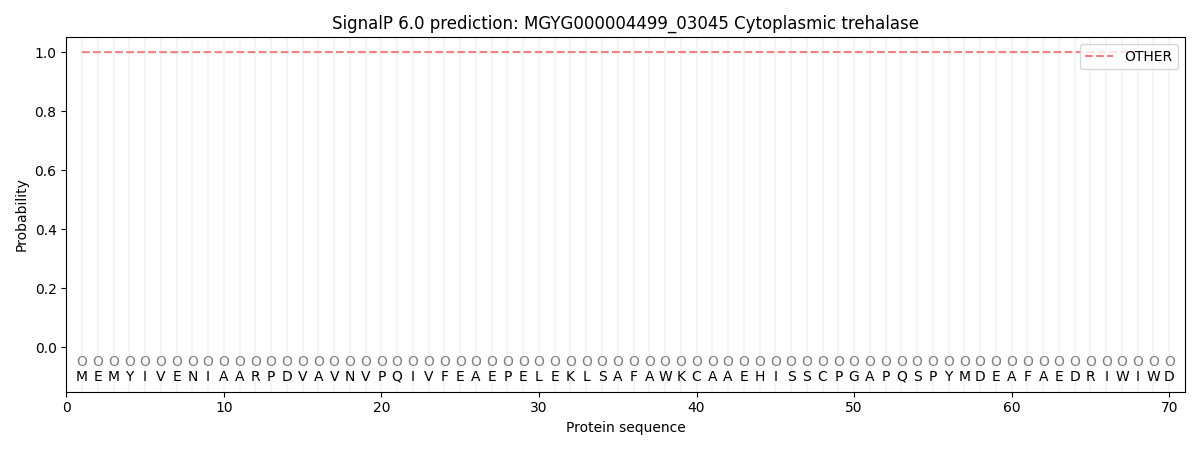
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999898 | 0.000149 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
