You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004510_01123
You are here: Home > Sequence: MGYG000004510_01123
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Prevotella sp002481295 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; Prevotella sp002481295 | |||||||||||
| CAZyme ID | MGYG000004510_01123 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | Undecaprenyl-phosphate 4-deoxy-4-formamido-L-arabinose transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 13868; End: 17266 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 776 | 1001 | 2.2e-37 | 0.991304347826087 |
| GH18 | 126 | 411 | 2.2e-34 | 0.8716216216216216 |
| CE4 | 497 | 599 | 2.5e-26 | 0.7769230769230769 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd10962 | CE4_GT2-like | 3.47e-91 | 499 | 697 | 1 | 196 | Catalytic NodB homology domain of uncharacterized bacterial glycosyl transferase, group 2-like family proteins. This family includes many uncharacterized bacterial proteins containing an N-terminal GH18 (glycosyl hydrolase, family 18) domain, a middle NodB-like homology domain, and a C-terminal GT2-like (glycosyl transferase group 2) domain. Although their biological function is unknown, members in this family contain a middle NodB homology domain that is similar to the catalytic domain of Streptococcus pneumoniae polysaccharide deacetylase PgdA (SpPgdA), an extracellular metal-dependent polysaccharide deacetylase with de-N-acetylase activity toward a hexamer of chitooligosaccharide N-acetylglucosamine, but not shorter chitooligosaccharides or a synthetic peptidoglycan tetrasaccharide. Like SpPgdA, this family is a member of the carbohydrate esterase 4 (CE4) superfamily. The presence of three domains suggests that members of this family may be multifunctional. |
| cd06423 | CESA_like | 2.41e-66 | 780 | 958 | 1 | 180 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| COG1215 | BcsA | 8.42e-63 | 725 | 1132 | 5 | 430 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| cd06549 | GH18_trifunctional | 1.68e-62 | 122 | 416 | 2 | 298 | GH18 domain of an uncharacterized family of bacterial proteins, which share a common three-domain architecture: an N-terminal glycosyl hydrolase family 18 (GH18) domain, a glycosyl transferase family 2 domain, and a C-terminal polysaccharide deacetylase domain. |
| PRK11204 | PRK11204 | 2.69e-61 | 720 | 1121 | 1 | 411 | N-glycosyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUB60716.1 | 0.0 | 1 | 1131 | 1 | 1132 |
| QUB69936.1 | 0.0 | 1 | 1131 | 1 | 1132 |
| QUB64534.1 | 0.0 | 1 | 1131 | 1 | 1132 |
| QUB59098.1 | 0.0 | 1 | 1131 | 1 | 1132 |
| QKH88098.1 | 0.0 | 1 | 1131 | 1 | 1132 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5O6Y_A | 3.33e-26 | 488 | 700 | 10 | 212 | Crystalstructure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579] |
| 7FBW_A | 1.44e-25 | 499 | 696 | 117 | 300 | ChainA, Predicted xylanase/chitin deacetylase [Caldanaerobacter subterraneus subsp. tengcongensis MB4] |
| 2C1G_A | 1.95e-25 | 499 | 697 | 236 | 417 | Structureof Streptococcus pneumoniae peptidoglycan deacetylase (SpPgdA) [Streptococcus pneumoniae R6] |
| 5JH8_A | 3.40e-25 | 207 | 424 | 89 | 315 | Crystalstructure of chitinase from Chromobacterium violaceum ATCC 12472 [Chromobacterium violaceum ATCC 12472] |
| 5O6Y_B | 3.89e-25 | 488 | 700 | 10 | 212 | Crystalstructure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579],5O6Y_C Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579],5O6Y_D Crystal structure of the Bc1960 peptidoglycan N-acetylglucosamine deacetylase in complex with 4-naphthalen-1-yl-~{N}-oxidanyl-benzamide [Bacillus cereus ATCC 14579] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8XAR5 | 1.52e-34 | 749 | 1038 | 48 | 330 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Escherichia coli O157:H7 OX=83334 GN=pgaC PE=3 SV=1 |
| P75905 | 5.02e-34 | 749 | 1038 | 48 | 330 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Escherichia coli (strain K12) OX=83333 GN=pgaC PE=1 SV=1 |
| Q6GDD8 | 3.28e-33 | 749 | 1036 | 21 | 299 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain MRSA252) OX=282458 GN=icaA PE=3 SV=1 |
| Q9RQP9 | 3.28e-33 | 749 | 1036 | 21 | 299 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=icaA PE=3 SV=2 |
| Q5HCN1 | 3.28e-33 | 749 | 1036 | 21 | 299 | Poly-beta-1,6-N-acetyl-D-glucosamine synthase OS=Staphylococcus aureus (strain COL) OX=93062 GN=icaA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
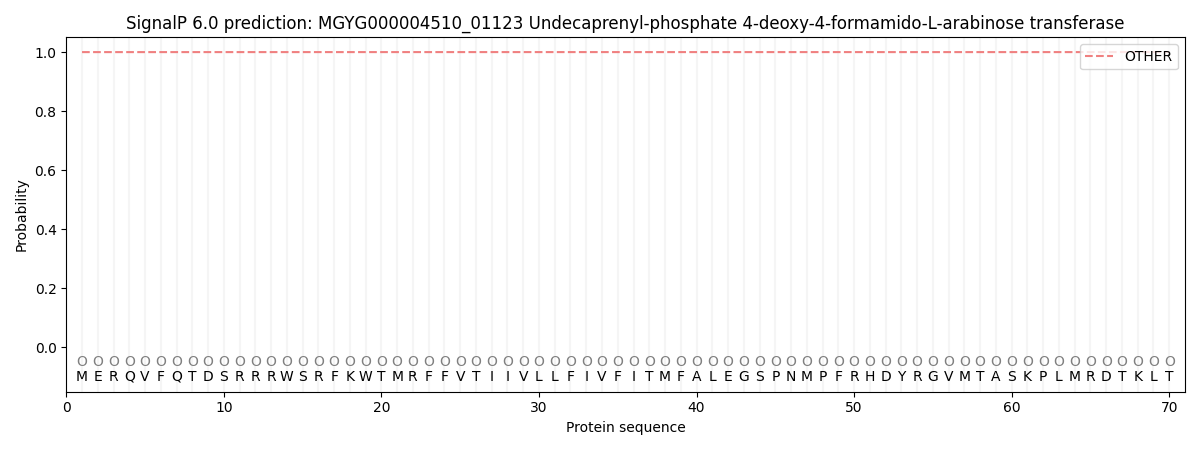
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000013 | 0.000002 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
TMHMM Annotations download full data without filtering help
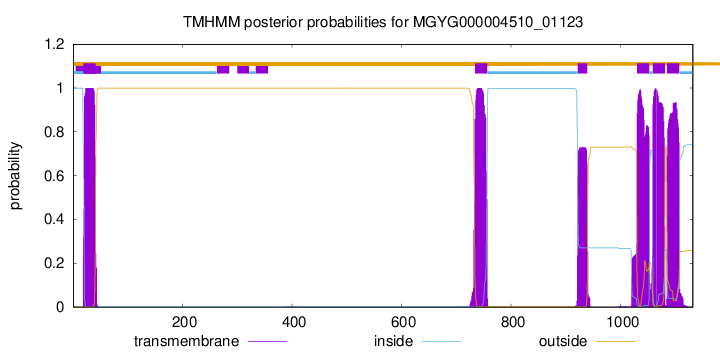
| start | end |
|---|---|
| 19 | 41 |
| 734 | 756 |
| 922 | 939 |
| 1030 | 1052 |
| 1059 | 1081 |
| 1085 | 1107 |
