You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004515_00239
You are here: Home > Sequence: MGYG000004515_00239
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
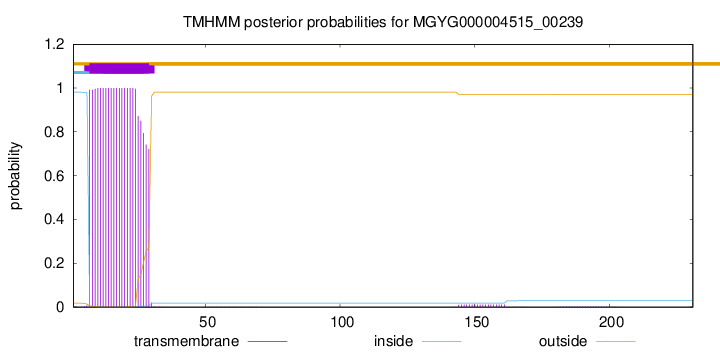
TMHMM annotations
Basic Information help
| Species | UMGS1663 sp900753245 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; TANB77; CAG-508; UMGS1663; UMGS1663 sp900753245 | |||||||||||
| CAZyme ID | MGYG000004515_00239 | |||||||||||
| CAZy Family | GT51 | |||||||||||
| CAZyme Description | Penicillin-binding protein 1F | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 95902; End: 96597 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT51 | 44 | 215 | 2.3e-61 | 0.9661016949152542 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00912 | Transgly | 9.73e-75 | 52 | 215 | 15 | 177 | Transglycosylase. The penicillin-binding proteins are bifunctional proteins consisting of transglycosylase and transpeptidase in the N- and C-terminus respectively. The transglycosylase domain catalyzes the polymerization of murein glycan chains. |
| COG0744 | MrcB | 5.34e-71 | 52 | 230 | 78 | 256 | Membrane carboxypeptidase (penicillin-binding protein) [Cell wall/membrane/envelope biogenesis]. |
| COG5009 | MrcA | 6.46e-62 | 43 | 231 | 59 | 247 | Membrane carboxypeptidase/penicillin-binding protein [Cell wall/membrane/envelope biogenesis]. |
| COG4953 | PbpC | 1.30e-40 | 53 | 225 | 61 | 231 | Membrane carboxypeptidase/penicillin-binding protein PbpC [Cell wall/membrane/envelope biogenesis]. |
| PRK11636 | mrcA | 5.67e-40 | 53 | 230 | 69 | 245 | penicillin-binding protein 1a; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUO38940.1 | 1.18e-79 | 1 | 229 | 1 | 232 |
| QNM09777.1 | 1.89e-73 | 6 | 229 | 19 | 241 |
| QRP41835.1 | 8.65e-73 | 22 | 225 | 25 | 227 |
| ASN93505.1 | 8.65e-73 | 22 | 225 | 25 | 227 |
| QJU22158.1 | 8.65e-73 | 22 | 225 | 25 | 227 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3NB6_A | 4.10e-45 | 52 | 231 | 22 | 200 | Crystalstructure of Aquifex aeolicus peptidoglycan glycosyltransferase in complex with Methylphosphoryl Neryl Moenomycin [Aquifex aeolicus] |
| 2OQO_A | 5.79e-45 | 52 | 231 | 22 | 200 | Crystalstructure of a peptidoglycan glycosyltransferase from a class A PBP: insight into bacterial cell wall synthesis [Aquifex aeolicus VF5],3D3H_A Crystal structure of a complex of the peptidoglycan glycosyltransferase domain from Aquifex aeolicus and neryl moenomycin A [Aquifex aeolicus],3NB7_A Crystal structure of Aquifex Aeolicus Peptidoglycan Glycosyltransferase in complex with Decarboxylated Neryl Moenomycin [Aquifex aeolicus] |
| 3UDF_A | 1.86e-38 | 53 | 229 | 42 | 217 | ChainA, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDF_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDI_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDI_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDX_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UDX_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE0_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE0_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE1_A Chain A, Penicillin-binding protein 1a [Acinetobacter baumannii],3UE1_B Chain B, Penicillin-binding protein 1a [Acinetobacter baumannii] |
| 4OON_A | 2.95e-38 | 57 | 223 | 46 | 211 | Crystalstructure of PBP1a in complex with compound 17 ((4Z,8S,11E,14S)-5-(2-amino-1,3-thiazol-4-yl)-14-(5,6-dihydroxy-1,3-dioxo-1,3-dihydro-2H-isoindol-2-yl)-8-formyl-2-methyl-6-oxo-3,10-dioxa-4,7,11-triazatetradeca-4,11-diene-2,12,14-tricarboxylic acid) [Pseudomonas aeruginosa PAO1] |
| 3DWK_A | 2.41e-32 | 51 | 214 | 28 | 190 | ChainA, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL],3DWK_B Chain B, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL],3DWK_C Chain C, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL],3DWK_D Chain D, Penicillin-binding protein 2 [Staphylococcus aureus subsp. aureus COL] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P38050 | 5.18e-47 | 41 | 225 | 57 | 241 | Penicillin-binding protein 1F OS=Bacillus subtilis (strain 168) OX=224308 GN=pbpF PE=2 SV=2 |
| O87579 | 7.45e-42 | 50 | 231 | 68 | 248 | Penicillin-binding protein 1A OS=Neisseria lactamica OX=486 GN=mrcA PE=3 SV=1 |
| Q07259 | 1.36e-41 | 41 | 229 | 38 | 224 | Putative transglycosylase H16_A0665 OS=Cupriavidus necator (strain ATCC 17699 / DSM 428 / KCTC 22496 / NCIMB 10442 / H16 / Stanier 337) OX=381666 GN=H16_A0665 PE=3 SV=2 |
| O87626 | 1.91e-41 | 50 | 231 | 68 | 248 | Penicillin-binding protein 1A OS=Neisseria flavescens OX=484 GN=mrcA PE=3 SV=1 |
| P0A0Z6 | 4.88e-41 | 50 | 231 | 68 | 248 | Penicillin-binding protein 1A OS=Neisseria meningitidis serogroup B (strain MC58) OX=122586 GN=mrcA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.928449 | 0.063015 | 0.006129 | 0.000321 | 0.000168 | 0.001937 |