You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004667_04414
You are here: Home > Sequence: MGYG000004667_04414
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium beijerinckii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium; Clostridium beijerinckii | |||||||||||
| CAZyme ID | MGYG000004667_04414 | |||||||||||
| CAZy Family | GH19 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 84; End: 1013 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH19 | 171 | 287 | 4.3e-18 | 0.5974025974025974 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3179 | COG3179 | 7.96e-16 | 134 | 297 | 11 | 197 | Predicted chitinase [General function prediction only]. |
| pfam01471 | PG_binding_1 | 2.27e-12 | 67 | 119 | 4 | 57 | Putative peptidoglycan binding domain. This domain is composed of three alpha helices. This domain is found at the N or C-terminus of a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. This family is found N-terminal to the catalytic domain of matrixins. The domain is found to bind peptidoglycan experimentally. |
| COG3409 | PGRP | 2.49e-12 | 3 | 119 | 39 | 183 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| COG3409 | PGRP | 3.37e-08 | 67 | 129 | 47 | 112 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| pfam01471 | PG_binding_1 | 2.24e-06 | 8 | 64 | 1 | 57 | Putative peptidoglycan binding domain. This domain is composed of three alpha helices. This domain is found at the N or C-terminus of a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. This family is found N-terminal to the catalytic domain of matrixins. The domain is found to bind peptidoglycan experimentally. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CUU49179.1 | 1.68e-226 | 1 | 309 | 1 | 309 |
| AJH00412.1 | 2.11e-220 | 1 | 309 | 1 | 309 |
| QUN33684.1 | 2.87e-218 | 1 | 309 | 1 | 309 |
| AVK47411.1 | 8.23e-218 | 1 | 309 | 1 | 309 |
| AQS06140.1 | 1.66e-217 | 1 | 309 | 1 | 309 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4OK7_A | 5.64e-07 | 154 | 303 | 79 | 221 | ChainA, Endolysin [Salmonella phage SPN1S],4OK7_B Chain B, Endolysin [Salmonella phage SPN1S],4OK7_C Chain C, Endolysin [Salmonella phage SPN1S] |
| 5TV7_A | 8.37e-07 | 6 | 115 | 74 | 165 | ChainA, Putative peptidoglycan-binding/hydrolysing protein [Clostridioides difficile 630],5TV7_B Chain B, Putative peptidoglycan-binding/hydrolysing protein [Clostridioides difficile 630] |
| 3BKH_A | 8.22e-06 | 85 | 131 | 39 | 85 | ChainA, lytic transglycosylase [Pseudomonas phage phiKZ],3BKV_A Chain A, lytic transglycosylase [Pseudomonas phage phiKZ] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O64203 | 2.82e-16 | 171 | 303 | 217 | 360 | Endolysin A OS=Mycobacterium phage D29 OX=28369 GN=10 PE=1 SV=1 |
| P44187 | 7.45e-09 | 198 | 297 | 97 | 191 | Glycosyl hydrolase family 19 domain-containing protein HI_1415 OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=HI_1415 PE=3 SV=1 |
| L7N653 | 1.17e-07 | 33 | 119 | 52 | 160 | N-acetylmuramoyl-L-alanine amidase CwlM OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=cwlM PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
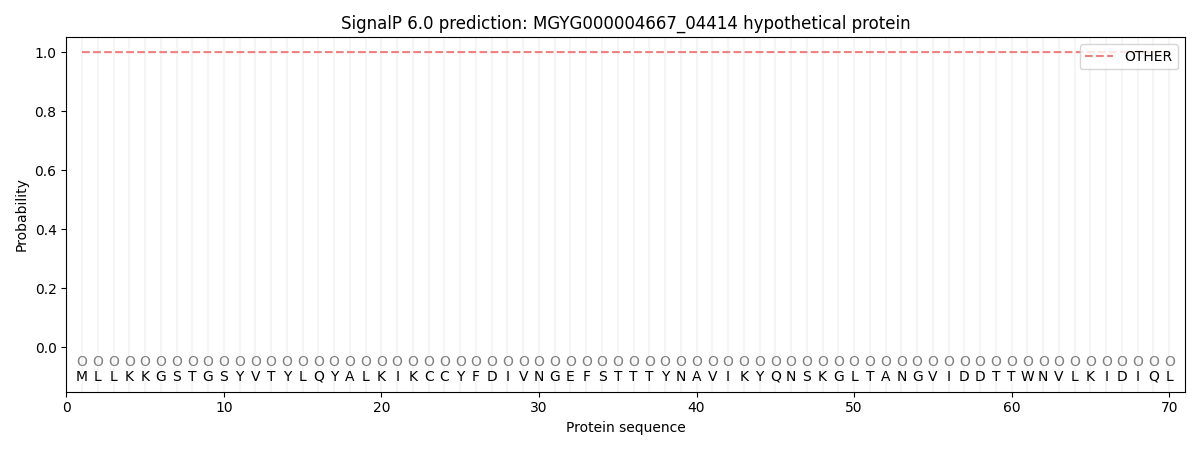
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999978 | 0.000076 | 0.000003 | 0.000000 | 0.000000 | 0.000000 |
