You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004697_02188
You are here: Home > Sequence: MGYG000004697_02188
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Eubacterium_F; | |||||||||||
| CAZyme ID | MGYG000004697_02188 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Teichoic acid poly(glycerol phosphate) polymerase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2941; End: 4125 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam04464 | Glyphos_transf | 2.02e-123 | 25 | 390 | 1 | 360 | CDP-Glycerol:Poly(glycerophosphate) glycerophosphotransferase. Wall-associated teichoic acids are a heterogeneous class of phosphate-rich polymers that are covalently linked to the cell wall peptidoglycan of gram-positive bacteria. They consist of a main chain of phosphodiester-linked polyols and/or sugar moieties attached to peptidoglycan via a linkage unit. CDP-glycerol:poly(glycerophosphate) glycerophosphotransferase is responsible for the polymerization of the main chain of the teichoic acid by sequential transfer of glycerol-phosphate units from CDP-glycerol to the linkage unit lipid. |
| COG1887 | TagB | 9.57e-88 | 1 | 391 | 7 | 387 | CDP-glycerol glycerophosphotransferase, TagB/SpsB family [Cell wall/membrane/envelope biogenesis, Lipid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQZ10349.1 | 5.70e-117 | 10 | 388 | 329 | 703 |
| QOV10415.1 | 2.44e-116 | 25 | 388 | 347 | 706 |
| QHS23522.1 | 8.37e-114 | 13 | 388 | 333 | 708 |
| CUH93585.1 | 4.00e-111 | 25 | 388 | 431 | 791 |
| AWC30558.1 | 1.07e-107 | 16 | 384 | 335 | 704 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3L7I_A | 1.77e-91 | 26 | 393 | 350 | 719 | Structureof the Wall Teichoic Acid Polymerase TagF [Staphylococcus epidermidis RP62A],3L7I_B Structure of the Wall Teichoic Acid Polymerase TagF [Staphylococcus epidermidis RP62A],3L7I_C Structure of the Wall Teichoic Acid Polymerase TagF [Staphylococcus epidermidis RP62A],3L7I_D Structure of the Wall Teichoic Acid Polymerase TagF [Staphylococcus epidermidis RP62A] |
| 3L7J_A | 1.90e-90 | 26 | 393 | 350 | 719 | ChainA, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7J_B Chain B, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7J_C Chain C, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7J_D Chain D, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7K_A Chain A, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7K_B Chain B, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7K_C Chain C, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7K_D Chain D, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7L_A Chain A, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7L_B Chain B, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7L_C Chain C, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7L_D Chain D, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A] |
| 3L7M_A | 5.25e-90 | 26 | 393 | 350 | 719 | ChainA, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7M_B Chain B, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7M_C Chain C, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A],3L7M_D Chain D, Teichoic acid biosynthesis protein F [Staphylococcus epidermidis RP62A] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8RKI5 | 2.11e-108 | 10 | 388 | 10 | 392 | Teichoic acid poly(glycerol phosphate) polymerase OS=Bacillus spizizenii (strain ATCC 23059 / NRRL B-14472 / W23) OX=655816 GN=tarF PE=1 SV=1 |
| P13485 | 2.40e-105 | 20 | 388 | 373 | 742 | Teichoic acid poly(glycerol phosphate) polymerase OS=Bacillus subtilis (strain 168) OX=224308 GN=tagF PE=1 SV=1 |
| Q2G1C1 | 1.76e-100 | 27 | 388 | 26 | 387 | Teichoic acid glycerol-phosphate transferase OS=Staphylococcus aureus (strain NCTC 8325 / PS 47) OX=93061 GN=tarF PE=1 SV=1 |
| Q5HLM5 | 8.18e-91 | 26 | 393 | 350 | 719 | Teichoic acid poly(glycerol phosphate) polymerase OS=Staphylococcus epidermidis (strain ATCC 35984 / RP62A) OX=176279 GN=tagF PE=1 SV=1 |
| P27621 | 3.64e-34 | 86 | 392 | 89 | 380 | Teichoic acid glycerol-phosphate primase OS=Bacillus subtilis (strain 168) OX=224308 GN=tagB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
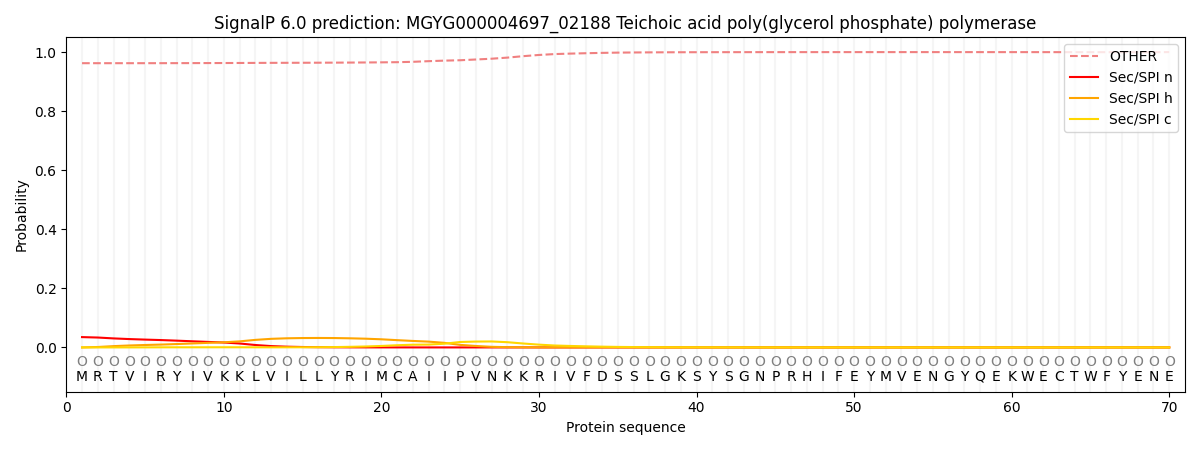
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.964742 | 0.032330 | 0.002706 | 0.000076 | 0.000044 | 0.000101 |
