You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004708_00135
You are here: Home > Sequence: MGYG000004708_00135
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Prevotella; | |||||||||||
| CAZyme ID | MGYG000004708_00135 | |||||||||||
| CAZy Family | CE19 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 22503; End: 23699 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE19 | 61 | 388 | 3e-24 | 0.8554216867469879 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0412 | DLH | 4.62e-14 | 117 | 270 | 21 | 141 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
| COG1506 | DAP2 | 2.27e-10 | 90 | 331 | 358 | 567 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| pfam12715 | Abhydrolase_7 | 1.01e-09 | 51 | 339 | 39 | 326 | Abhydrolase family. This is a family of probable bacterial abhydrolases. |
| pfam00326 | Peptidase_S9 | 1.19e-06 | 173 | 326 | 9 | 155 | Prolyl oligopeptidase family. |
| COG3458 | Axe1 | 0.006 | 223 | 271 | 157 | 206 | Cephalosporin-C deacetylase or related acetyl esterase [Secondary metabolites biosynthesis, transport and catabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADY37157.1 | 1.24e-133 | 32 | 379 | 379 | 726 |
| QEH40625.1 | 9.37e-20 | 65 | 382 | 51 | 359 |
| AMV40306.1 | 7.35e-18 | 70 | 384 | 69 | 365 |
| QNN25394.1 | 7.55e-16 | 61 | 387 | 51 | 362 |
| QEG26086.1 | 1.65e-15 | 61 | 381 | 26 | 329 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2XLC_A | 2.16e-07 | 58 | 270 | 26 | 202 | Acetylxylan esterase from Bacillus pumilus CECT5072 bound to paraoxon [Bacillus pumilus],2XLC_B Acetyl xylan esterase from Bacillus pumilus CECT5072 bound to paraoxon [Bacillus pumilus],2XLC_C Acetyl xylan esterase from Bacillus pumilus CECT5072 bound to paraoxon [Bacillus pumilus],2XLC_D Acetyl xylan esterase from Bacillus pumilus CECT5072 bound to paraoxon [Bacillus pumilus],2XLC_E Acetyl xylan esterase from Bacillus pumilus CECT5072 bound to paraoxon [Bacillus pumilus],2XLC_F Acetyl xylan esterase from Bacillus pumilus CECT5072 bound to paraoxon [Bacillus pumilus] |
| 2XLB_A | 6.81e-07 | 58 | 270 | 26 | 202 | Acetylxylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_B Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_C Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_D Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_E Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_F Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_G Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_H Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_I Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_J Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_K Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus],2XLB_L Acetyl xylan esterase from Bacillus pumilus without ligands [Bacillus pumilus] |
| 3FVR_A | 9.08e-07 | 58 | 270 | 26 | 202 | CrystalStructure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_B Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_C Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_D Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_E Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_F Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_G Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_H Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_I Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_L Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_M Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVR_N Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form I [Bacillus pumilus],3FVT_A Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_B Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_C Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_D Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_E Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_F Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_G Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_H Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_I Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_L Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_M Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FVT_N Crystal Structure of Acetyl Xylan Esterase from Bacillus pumilus, monoclinic crystal form II [Bacillus pumilus],3FYU_C Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_E Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_F Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_G Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_L Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_M Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_N Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus] |
| 3FYT_A | 2.15e-06 | 58 | 270 | 26 | 202 | ChainA, Acetyl xylan esterase [Bacillus pumilus],3FYT_B Chain B, Acetyl xylan esterase [Bacillus pumilus],3FYT_C Chain C, Acetyl xylan esterase [Bacillus pumilus],3FYT_D Chain D, Acetyl xylan esterase [Bacillus pumilus],3FYT_E Chain E, Acetyl xylan esterase [Bacillus pumilus],3FYT_F Chain F, Acetyl xylan esterase [Bacillus pumilus],3FYT_G Chain G, Acetyl xylan esterase [Bacillus pumilus],3FYT_H Chain H, Acetyl xylan esterase [Bacillus pumilus],3FYT_I Chain I, Acetyl xylan esterase [Bacillus pumilus],3FYT_L Chain L, Acetyl xylan esterase [Bacillus pumilus],3FYT_M Chain M, Acetyl xylan esterase [Bacillus pumilus],3FYT_N Chain N, Acetyl xylan esterase [Bacillus pumilus] |
| 3FYU_A | 3.81e-06 | 58 | 270 | 26 | 202 | Crystalstructure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_B Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_D Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_H Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus],3FYU_I Crystal structure of acetyl xylan esterase from Bacillus pumilus obtained in presence of D-xylose and sodium acetate [Bacillus pumilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34973 | 8.86e-26 | 63 | 381 | 3 | 299 | Putative hydrolase YtaP OS=Bacillus subtilis (strain 168) OX=224308 GN=ytaP PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
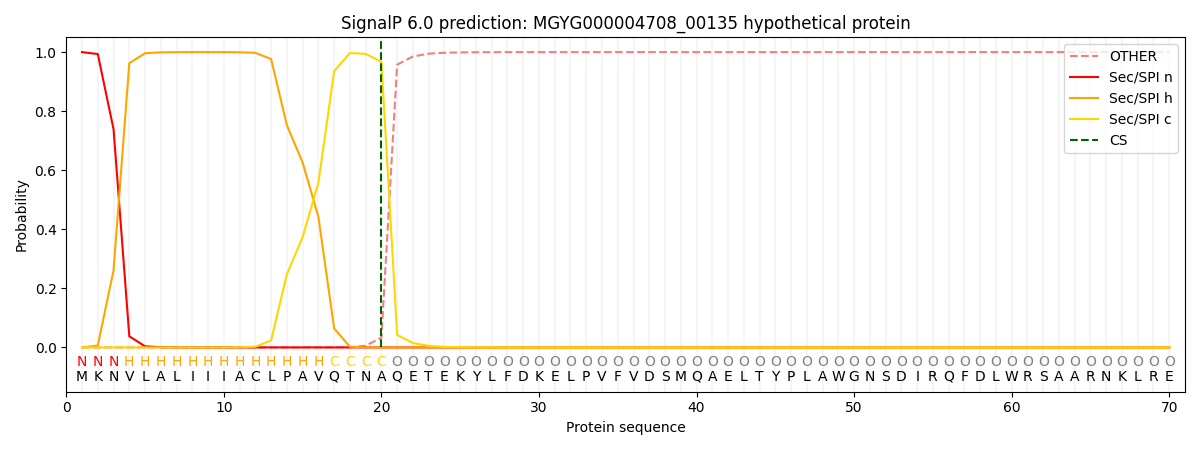
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000307 | 0.998833 | 0.000341 | 0.000182 | 0.000158 | 0.000143 |
