You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004720_01836
You are here: Home > Sequence: MGYG000004720_01836
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paramuribaculum sp900759915 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Muribaculaceae; Paramuribaculum; Paramuribaculum sp900759915 | |||||||||||
| CAZyme ID | MGYG000004720_01836 | |||||||||||
| CAZy Family | GH65 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 12616; End: 14592 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH65 | 291 | 626 | 1.5e-88 | 0.9865591397849462 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1554 | ATH1 | 4.54e-69 | 42 | 633 | 31 | 718 | Trehalose and maltose hydrolase (possible phosphorylase) [Carbohydrate transport and metabolism]. |
| pfam03632 | Glyco_hydro_65m | 3.41e-46 | 291 | 622 | 3 | 379 | Glycosyl hydrolase family 65 central catalytic domain. This family of glycosyl hydrolases contains vacuolar acid trehalase and maltose phosphorylase.Maltose phosphorylase (MP) is a dimeric enzyme that catalyzes the conversion of maltose and inorganic phosphate into beta-D-glucose-1-phosphate and glucose. The central domain is the catalytic domain, which binds a phosphate ion that is proximal the the highly conserved Glu. The arrangement of the phosphate and the glutamate is thought to cause nucleophilic attack on the anomeric carbon atom. The catalytic domain also forms the majority of the dimerization interface. |
| PRK13807 | PRK13807 | 3.53e-17 | 218 | 474 | 247 | 507 | maltose phosphorylase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBK63836.1 | 6.63e-191 | 11 | 658 | 10 | 664 |
| AFL77434.1 | 9.38e-191 | 17 | 658 | 16 | 664 |
| QUT91743.1 | 1.57e-169 | 38 | 646 | 38 | 660 |
| ADY37608.1 | 1.14e-164 | 28 | 647 | 29 | 661 |
| QRQ48989.1 | 1.20e-162 | 38 | 646 | 36 | 657 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7FE3_A | 3.52e-133 | 35 | 647 | 33 | 665 | ChainA, Candidate alpha glycoside phosphorylase Glycoside hydrolase family 65 [Flavobacterium johnsoniae UW101],7FE3_B Chain B, Candidate alpha glycoside phosphorylase Glycoside hydrolase family 65 [Flavobacterium johnsoniae UW101],7FE3_C Chain C, Candidate alpha glycoside phosphorylase Glycoside hydrolase family 65 [Flavobacterium johnsoniae UW101],7FE4_A Chain A, Candidate alpha glycoside phosphorylase Glycoside hydrolase family 65 [Flavobacterium johnsoniae UW101],7FE4_B Chain B, Candidate alpha glycoside phosphorylase Glycoside hydrolase family 65 [Flavobacterium johnsoniae UW101],7FE4_C Chain C, Candidate alpha glycoside phosphorylase Glycoside hydrolase family 65 [Flavobacterium johnsoniae UW101] |
| 3WIQ_A | 1.33e-41 | 205 | 644 | 223 | 714 | Crystalstructure of kojibiose phosphorylase complexed with kojibiose [Caldicellulosiruptor saccharolyticus DSM 8903],3WIR_A Crystal structure of kojibiose phosphorylase complexed with glucose [Caldicellulosiruptor saccharolyticus DSM 8903],3WIR_B Crystal structure of kojibiose phosphorylase complexed with glucose [Caldicellulosiruptor saccharolyticus DSM 8903],3WIR_C Crystal structure of kojibiose phosphorylase complexed with glucose [Caldicellulosiruptor saccharolyticus DSM 8903],3WIR_D Crystal structure of kojibiose phosphorylase complexed with glucose [Caldicellulosiruptor saccharolyticus DSM 8903] |
| 4KTP_A | 2.64e-25 | 218 | 636 | 229 | 705 | ChainA, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTP_B Chain B, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10] |
| 4KTR_A | 6.15e-25 | 218 | 636 | 229 | 705 | ChainA, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_B Chain B, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_C Chain C, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_D Chain D, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_E Chain E, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_F Chain F, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_G Chain G, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10],4KTR_H Chain H, Glycoside hydrolase family 65 central catalytic [[Bacillus] selenitireducens MLS10] |
| 1H54_A | 5.37e-23 | 207 | 640 | 233 | 705 | ChainA, Maltose Phosphorylase [Levilactobacillus brevis],1H54_B Chain B, Maltose Phosphorylase [Levilactobacillus brevis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q54KX5 | 3.51e-70 | 26 | 633 | 25 | 645 | Protein-glucosylgalactosylhydroxylysine glucosidase OS=Dictyostelium discoideum OX=44689 GN=pgghg PE=3 SV=2 |
| F1NZI4 | 1.93e-59 | 253 | 647 | 230 | 635 | Protein-glucosylgalactosylhydroxylysine glucosidase OS=Gallus gallus OX=9031 GN=PGGHG PE=1 SV=3 |
| Q32M88 | 7.70e-58 | 218 | 658 | 187 | 633 | Protein-glucosylgalactosylhydroxylysine glucosidase OS=Homo sapiens OX=9606 GN=PGGHG PE=1 SV=2 |
| Q8BP56 | 5.98e-55 | 243 | 634 | 206 | 612 | Protein-glucosylgalactosylhydroxylysine glucosidase OS=Mus musculus OX=10090 GN=Pgghg PE=1 SV=1 |
| A0JMP0 | 1.28e-54 | 248 | 646 | 167 | 582 | Protein-glucosylgalactosylhydroxylysine glucosidase OS=Danio rerio OX=7955 GN=pgghg PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
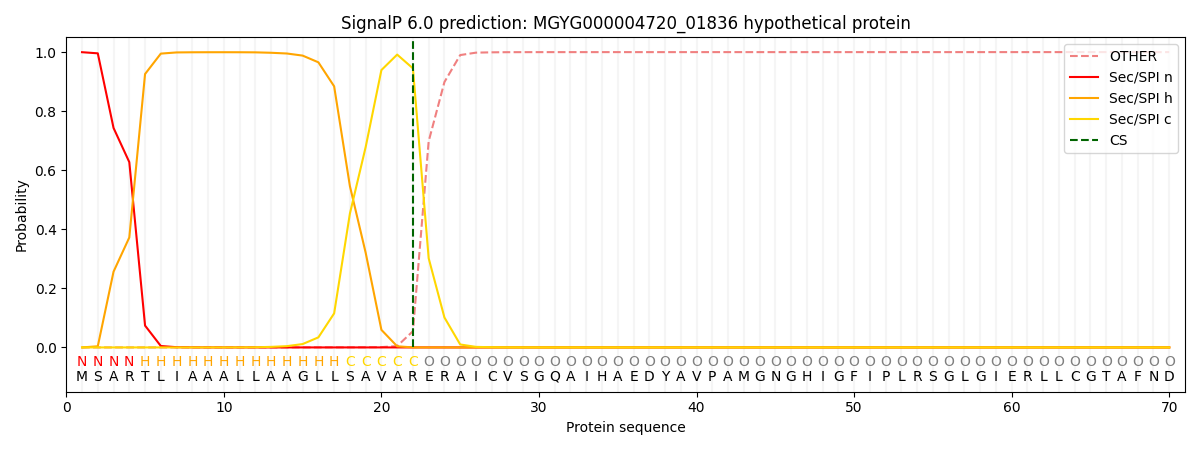
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000570 | 0.998070 | 0.000755 | 0.000207 | 0.000181 | 0.000174 |
