You are browsing environment: HUMAN GUT
MGYG000004727_00332
Basic Information
help
Species
Lineage
Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; ;
CAZyme ID
MGYG000004727_00332
CAZy Family
GH55
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000004727
2194019
MAG
China
Asia
Gene Location
Start: 4331;
End: 7279
Strand: -
No EC number prediction in MGYG000004727_00332.
Family
Start
End
Evalue
family coverage
GH55
36
605
2.5e-74
0.7837837837837838
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam12708
Pectate_lyase_3
1.60e-13
367
537
1
213
Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28.
more
COG5434
Pgu1
9.99e-05
369
498
84
211
Polygalacturonase [Carbohydrate transport and metabolism].
more
COG3386
YvrE
0.004
673
761
129
231
Sugar lactone lactonase YvrE [Carbohydrate transport and metabolism].
more
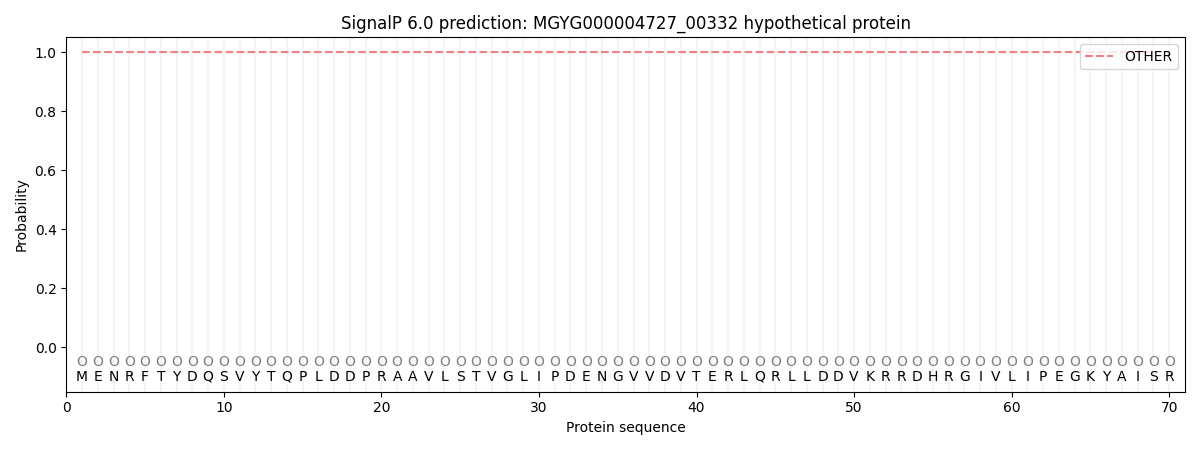
This protein is predicted as OTHER
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
1.000060
0.000000
0.000000
0.000000
0.000000
0.000000
There is no transmembrane helices in MGYG000004727_00332.