You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004799_01937
You are here: Home > Sequence: MGYG000004799_01937
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Schaedlerella sp900765975 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Schaedlerella; Schaedlerella sp900765975 | |||||||||||
| CAZyme ID | MGYG000004799_01937 | |||||||||||
| CAZy Family | GH1 | |||||||||||
| CAZyme Description | 1,4-beta-D-glucan glucohydrolase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 10518; End: 11804 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH1 | 2 | 424 | 1.9e-116 | 0.972027972027972 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG2723 | BglB | 2.91e-91 | 1 | 422 | 1 | 444 | Beta-glucosidase/6-phospho-beta-glucosidase/beta-galactosidase [Carbohydrate transport and metabolism]. |
| TIGR03356 | BGL | 3.99e-84 | 7 | 423 | 3 | 425 | beta-galactosidase. |
| pfam00232 | Glyco_hydro_1 | 1.64e-72 | 4 | 424 | 5 | 444 | Glycosyl hydrolase family 1. |
| PRK13511 | PRK13511 | 2.14e-51 | 1 | 423 | 2 | 458 | 6-phospho-beta-galactosidase; Provisional |
| PLN02814 | PLN02814 | 1.70e-33 | 4 | 425 | 28 | 478 | beta-glucosidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEN97245.1 | 5.49e-222 | 8 | 426 | 7 | 431 |
| QPK81646.1 | 3.66e-204 | 1 | 426 | 1 | 425 |
| QTE67958.1 | 9.07e-204 | 4 | 425 | 3 | 423 |
| AXB29242.1 | 4.18e-200 | 1 | 426 | 1 | 429 |
| CBL02685.1 | 6.50e-197 | 1 | 425 | 1 | 428 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4R27_A | 1.45e-114 | 4 | 426 | 7 | 407 | Crystalstructure of beta-glycosidase BGL167 [Microbacterium sp. Gsoil167],4R27_B Crystal structure of beta-glycosidase BGL167 [Microbacterium sp. Gsoil167] |
| 6IER_A | 5.68e-94 | 6 | 425 | 34 | 425 | Apostructure of a beta-glucosidase 1317 [uncultured bacterium] |
| 6Z1H_A | 4.21e-64 | 3 | 423 | 10 | 440 | ChainA, ANCESTRAL RECONSTRUCTED GLYCOSIDASE [synthetic construct],6Z1H_B Chain B, ANCESTRAL RECONSTRUCTED GLYCOSIDASE [synthetic construct],6Z1M_A Chain A, Ancestral reconstructed glycosidase [synthetic construct],6Z1M_B Chain B, Ancestral reconstructed glycosidase [synthetic construct],6Z1M_C Chain C, Ancestral reconstructed glycosidase [synthetic construct] |
| 1VFF_A | 4.77e-64 | 3 | 425 | 4 | 400 | beta-glycosidasefrom Pyrococcus horikoshii [Pyrococcus horikoshii] |
| 6ZIV_AAA | 4.09e-52 | 2 | 423 | 13 | 445 | ChainAAA, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_BBB Chain BBB, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_CCC Chain CCC, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_DDD Chain DDD, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_EEE Chain EEE, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_FFF Chain FFF, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_GGG Chain GGG, Beta-glucosidase [Alicyclobacillus tengchongensis],6ZIV_HHH Chain HHH, Beta-glucosidase [Alicyclobacillus tengchongensis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B9K7M5 | 2.95e-52 | 1 | 426 | 1 | 435 | 1,4-beta-D-glucan glucohydrolase OS=Thermotoga neapolitana (strain ATCC 49049 / DSM 4359 / NBRC 107923 / NS-E) OX=309803 GN=gghA PE=1 SV=2 |
| Q08638 | 2.36e-50 | 1 | 426 | 3 | 437 | Beta-glucosidase A OS=Thermotoga maritima (strain ATCC 43589 / DSM 3109 / JCM 10099 / NBRC 100826 / MSB8) OX=243274 GN=bglA PE=1 SV=1 |
| P0C946 | 5.91e-49 | 1 | 416 | 1 | 425 | 1,4-beta-D-glucan glucohydrolase (Fragment) OS=Thermotoga neapolitana OX=2337 GN=bglA PE=3 SV=1 |
| P10482 | 7.95e-49 | 4 | 423 | 5 | 446 | Beta-glucosidase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=bglA PE=3 SV=1 |
| P22505 | 3.76e-46 | 4 | 420 | 8 | 435 | Beta-glucosidase B OS=Paenibacillus polymyxa OX=1406 GN=bglB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
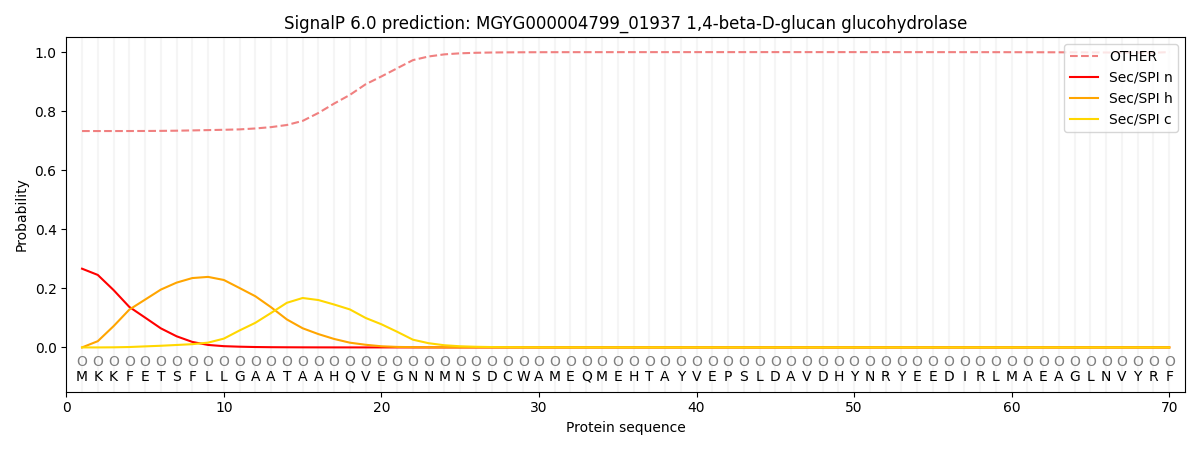
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.749870 | 0.248189 | 0.001057 | 0.000355 | 0.000210 | 0.000325 |
