You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004839_02494
Basic Information
help
| Species |
|
| Lineage |
Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; GCA-900066135;
|
| CAZyme ID |
MGYG000004839_02494
|
| CAZy Family |
CE19 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000004839 |
3024091 |
MAG |
China |
Asia |
|
| Gene Location |
Start: 6036;
End: 7127
Strand: -
|
No EC number prediction in MGYG000004839_02494.
| Family |
Start |
End |
Evalue |
family coverage |
| CE19 |
29 |
345 |
4.6e-33 |
0.8192771084337349 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam12715
|
Abhydrolase_7 |
7.70e-07 |
9 |
337 |
29 |
340 |
Abhydrolase family. This is a family of probable bacterial abhydrolases. |
| COG1506
|
DAP2 |
1.83e-06 |
69 |
288 |
360 |
537 |
Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| pfam00561
|
Abhydrolase_1 |
4.89e-04 |
134 |
246 |
13 |
97 |
alpha/beta hydrolase fold. This catalytic domain is found in a very wide range of enzymes. |
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
|
O34973
|
7.72e-16 |
37 |
331 |
2 |
275 |
Putative hydrolase YtaP OS=Bacillus subtilis (strain 168) OX=224308 GN=ytaP PE=3 SV=1 |
This protein is predicted as OTHER
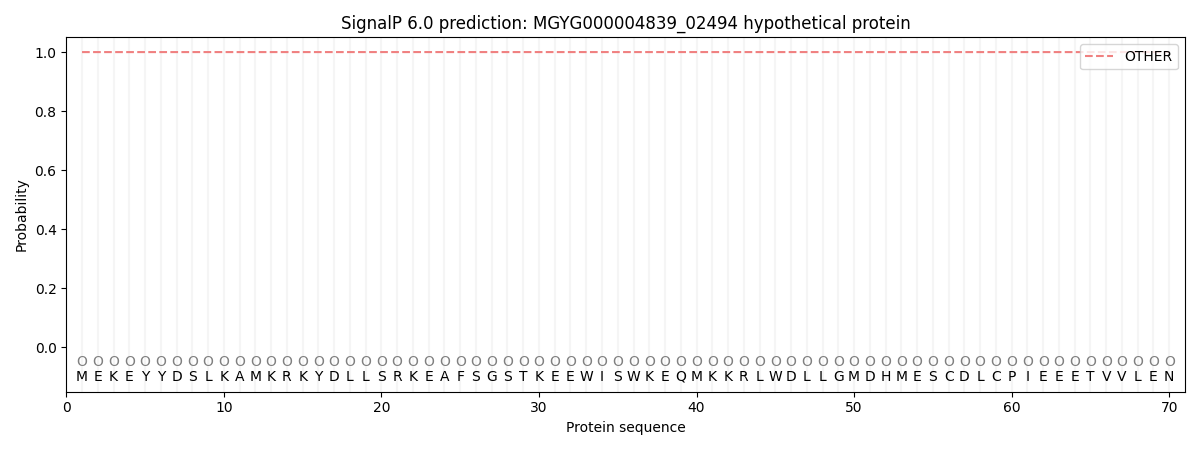
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000068
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000004839_02494.

