You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004859_00683
You are here: Home > Sequence: MGYG000004859_00683
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Collinsella sp008014645 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Coriobacteriia; Coriobacteriales; Coriobacteriaceae; Collinsella; Collinsella sp008014645 | |||||||||||
| CAZyme ID | MGYG000004859_00683 | |||||||||||
| CAZy Family | GH84 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9155; End: 13093 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH84 | 182 | 478 | 8.5e-97 | 0.9898305084745763 |
| CBM32 | 715 | 843 | 3.4e-24 | 0.9435483870967742 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam07555 | NAGidase | 2.65e-125 | 182 | 478 | 1 | 293 | beta-N-acetylglucosaminidase. This family has previously been described as a hyaluronidase. However, more recently it has been shown that this family has beta-N-acetylglucosaminidase activity. |
| pfam02838 | Glyco_hydro_20b | 3.03e-19 | 34 | 175 | 1 | 123 | Glycosyl hydrolase family 20, domain 2. This domain has a zincin-like fold. |
| pfam00754 | F5_F8_type_C | 2.14e-16 | 729 | 842 | 15 | 127 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| smart00089 | PKD | 2.32e-12 | 621 | 690 | 2 | 71 | Repeats in polycystic kidney disease 1 (PKD1) and other proteins. Polycystic kidney disease 1 protein contains 14 repeats, present elsewhere such as in microbial collagenases. |
| NF033190 | inl_like_NEAT_1 | 5.76e-10 | 1116 | 1309 | 575 | 754 | NEAT domain-containing leucine-rich repeat protein. Members of this family have an N-terminal NEAT (near transporter) domain often associated with iron transport, followed by a leucine-rich repeat region with significant sequence similarity to the internalins of Listeria monocytogenes. However, since Bacillus cereus (from which this protein was described, in PMID:16978259) is not considered an intracellular pathogen, and the function may be iron transport rather than internalization, applying the name "internalin" to this family probably would be misleading. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCJ44459.1 | 2.99e-210 | 36 | 924 | 40 | 934 |
| AZU64318.1 | 5.36e-203 | 37 | 924 | 42 | 935 |
| QZO76584.1 | 1.22e-146 | 151 | 856 | 819 | 1520 |
| QUY64130.1 | 4.34e-146 | 151 | 856 | 819 | 1520 |
| AOY53993.1 | 4.81e-117 | 35 | 617 | 44 | 621 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2CBI_A | 5.04e-123 | 35 | 617 | 14 | 591 | Structureof the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase [Clostridium perfringens],2CBI_B Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase [Clostridium perfringens],2CBJ_A Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc [Clostridium perfringens],2CBJ_B Structure of the Clostridium perfringens NagJ family 84 glycoside hydrolase, a homologue of human O-GlcNAcase in complex with PUGNAc [Clostridium perfringens],2V5C_A Family 84 glycoside hydrolase from Clostridium perfringens, 2.1 Angstrom structure [Clostridium perfringens],2V5C_B Family 84 glycoside hydrolase from Clostridium perfringens, 2.1 Angstrom structure [Clostridium perfringens],2VUR_A Chemical dissection of the link between Streptozotocin, O-GlcNAc and pancreatic cell death [Clostridium perfringens],2VUR_B Chemical dissection of the link between Streptozotocin, O-GlcNAc and pancreatic cell death [Clostridium perfringens],2X0Y_A Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds [Clostridium perfringens],2X0Y_B Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds [Clostridium perfringens] |
| 2J62_A | 7.01e-123 | 35 | 617 | 14 | 591 | Structureof a bacterial O-glcnacase in complex with glcnacstatin [Clostridium perfringens],2J62_B Structure of a bacterial O-glcnacase in complex with glcnacstatin [Clostridium perfringens],2WB5_A GlcNAcstatins are nanomolar inhibitors of human O-GlcNAcase inducing cellular hyper-O-GlcNAcylation [Clostridium perfringens],2WB5_B GlcNAcstatins are nanomolar inhibitors of human O-GlcNAcase inducing cellular hyper-O-GlcNAcylation [Clostridium perfringens] |
| 5OXD_A | 1.20e-122 | 35 | 605 | 16 | 566 | Complexof a C. perfringens O-GlcNAcase with a fragment hit [Clostridium perfringens] |
| 2XPK_A | 3.65e-122 | 35 | 617 | 14 | 591 | Cell-penetrant,nanomolar O-GlcNAcase inhibitors selective against lysosomal hexosaminidases [Clostridium perfringens],2XPK_B Cell-penetrant, nanomolar O-GlcNAcase inhibitors selective against lysosomal hexosaminidases [Clostridium perfringens] |
| 4ZXL_A | 4.50e-122 | 35 | 605 | 6 | 556 | CpOGAD298N in complex with Drosophila HCF -derived Thr-O-GlcNAc peptide [Clostridium perfringens ATCC 13124] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0TR53 | 9.63e-118 | 35 | 617 | 44 | 621 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain ATCC 13124 / DSM 756 / JCM 1290 / NCIMB 6125 / NCTC 8237 / Type A) OX=195103 GN=nagJ PE=1 SV=1 |
| Q8XL08 | 9.23e-117 | 35 | 617 | 44 | 621 | O-GlcNAcase NagJ OS=Clostridium perfringens (strain 13 / Type A) OX=195102 GN=nagJ PE=1 SV=1 |
| Q89ZI2 | 4.28e-68 | 128 | 510 | 99 | 474 | O-GlcNAcase BT_4395 OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_4395 PE=1 SV=1 |
| O60502 | 3.04e-33 | 183 | 485 | 63 | 378 | Protein O-GlcNAcase OS=Homo sapiens OX=9606 GN=OGA PE=1 SV=2 |
| Q8VIJ5 | 3.04e-33 | 183 | 485 | 63 | 378 | Protein O-GlcNAcase OS=Rattus norvegicus OX=10116 GN=Oga PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
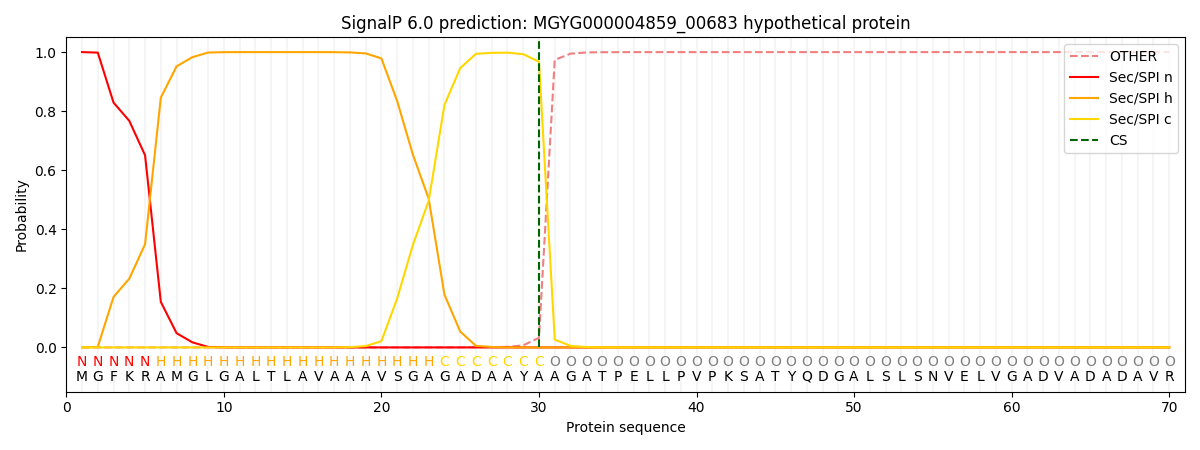
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000289 | 0.998958 | 0.000195 | 0.000206 | 0.000179 | 0.000153 |
