You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004861_00651
You are here: Home > Sequence: MGYG000004861_00651
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Limosilactobacillus sp012843675 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Limosilactobacillus; Limosilactobacillus sp012843675 | |||||||||||
| CAZyme ID | MGYG000004861_00651 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 44967; End: 46532 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00877 | NLPC_P60 | 1.02e-26 | 418 | 512 | 3 | 96 | NlpC/P60 family. The function of this domain is unknown. It is found in several lipoproteins. |
| COG0791 | Spr | 1.91e-16 | 418 | 510 | 89 | 185 | Cell wall-associated hydrolase, NlpC family [Cell wall/membrane/envelope biogenesis]. |
| PRK13914 | PRK13914 | 6.44e-15 | 21 | 499 | 14 | 460 | invasion associated endopeptidase. |
| cd00118 | LysM | 2.21e-14 | 101 | 144 | 2 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| smart00257 | LysM | 1.17e-13 | 102 | 144 | 2 | 44 | Lysin motif. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QOL69049.1 | 1.99e-136 | 1 | 521 | 1 | 513 |
| QLI94011.1 | 6.36e-135 | 1 | 521 | 1 | 513 |
| AJT51486.1 | 3.59e-133 | 5 | 521 | 1 | 509 |
| QLQ62351.1 | 2.04e-77 | 3 | 519 | 4 | 502 |
| QIZ04030.1 | 5.63e-77 | 3 | 519 | 4 | 502 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6B8C_A | 3.74e-25 | 418 | 519 | 42 | 142 | Crystalstructure of NlpC/p60 domain of peptidoglycan hydrolase SagA [Enterococcus faecium] |
| 4Q4T_A | 1.19e-11 | 412 | 515 | 351 | 465 | Structureof the Resuscitation Promoting Factor Interacting protein RipA mutated at E444 [Mycobacterium tuberculosis H37Rv] |
| 2K1G_A | 1.99e-08 | 417 | 495 | 19 | 97 | SolutionNMR structure of lipoprotein spr from Escherichia coli K12. Northeast Structural Genomics target ER541-37-162 [Escherichia coli K-12] |
| 3I86_A | 5.69e-08 | 417 | 515 | 25 | 134 | Crystalstructure of the P60 Domain from M. avium subspecies paratuberculosis antigen MAP1204 [Mycobacterium avium subsp. paratuberculosis],3I86_B Crystal structure of the P60 Domain from M. avium subspecies paratuberculosis antigen MAP1204 [Mycobacterium avium subsp. paratuberculosis] |
| 3H41_A | 1.34e-06 | 417 | 517 | 202 | 305 | CRYSTALSTRUCTURE OF A NLPC/P60 FAMILY PROTEIN (BCE_2878) FROM BACILLUS CEREUS ATCC 10987 AT 1.79 A RESOLUTION [Bacillus cereus ATCC 10987] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P13692 | 3.73e-22 | 418 | 519 | 416 | 516 | Protein P54 OS=Enterococcus faecium OX=1352 PE=3 SV=2 |
| O34669 | 2.45e-14 | 21 | 161 | 11 | 138 | Cell wall-binding protein YocH OS=Bacillus subtilis (strain 168) OX=224308 GN=yocH PE=1 SV=1 |
| O31852 | 1.73e-11 | 25 | 151 | 17 | 138 | D-gamma-glutamyl-meso-diaminopimelic acid endopeptidase CwlS OS=Bacillus subtilis (strain 168) OX=224308 GN=cwlS PE=1 SV=1 |
| A2RHZ5 | 2.50e-11 | 38 | 151 | 246 | 369 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
| P0C2T5 | 5.87e-11 | 38 | 151 | 246 | 369 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris OX=1359 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
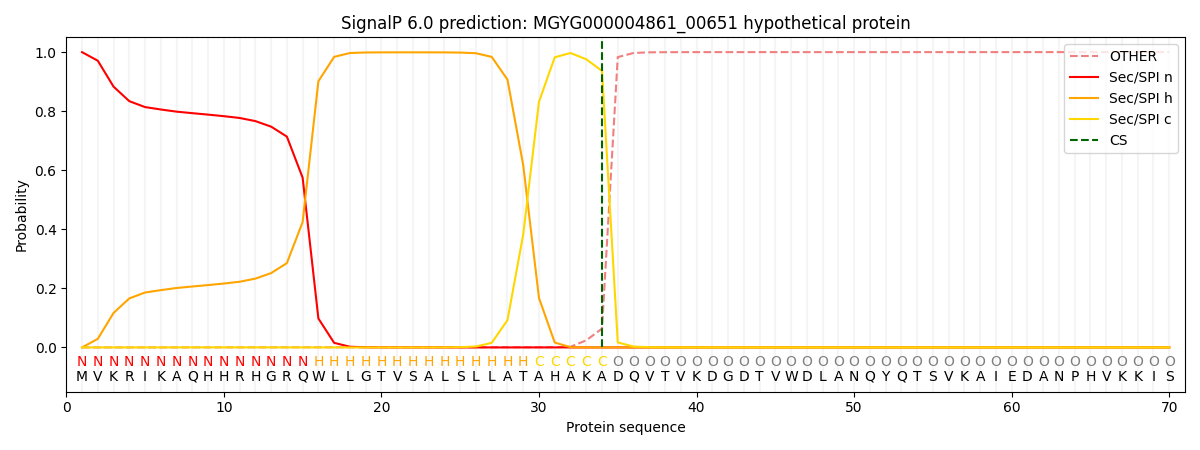
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000477 | 0.998502 | 0.000339 | 0.000276 | 0.000192 | 0.000159 |
