You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004899_03364
You are here: Home > Sequence: MGYG000004899_03364
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp900555635 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp900555635 | |||||||||||
| CAZyme ID | MGYG000004899_03364 | |||||||||||
| CAZy Family | GH55 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 61380; End: 64376 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH55 | 29 | 668 | 1.1e-90 | 0.8594594594594595 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3386 | YvrE | 4.72e-14 | 698 | 988 | 5 | 274 | Sugar lactone lactonase YvrE [Carbohydrate transport and metabolism]. |
| pfam12708 | Pectate_lyase_3 | 3.72e-12 | 36 | 208 | 7 | 171 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| pfam08450 | SGL | 9.99e-12 | 899 | 987 | 145 | 243 | SMP-30/Gluconolaconase/LRE-like region. This family describes a region that is found in proteins expressed by a variety of eukaryotic and prokaryotic species. These proteins include various enzymes, such as senescence marker protein 30 (SMP-30), gluconolactonase and luciferin-regenerating enzyme (LRE). SMP-30 is known to hydrolyze diisopropyl phosphorofluoridate in the liver, and has been noted as having sequence similarity, in the region described in this family, with PON1 and LRE. |
| pfam12708 | Pectate_lyase_3 | 4.14e-11 | 369 | 422 | 1 | 60 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| cd05819 | NHL | 8.61e-07 | 724 | 959 | 12 | 226 | NHL repeat unit of beta-propeller proteins. The NHL(NCL-1, HT2A and LIN-41)-repeat is found in multiple tandem copies, typically as 6 instances. It is about 40 residues long and resembles the WD repeat and other beta-propeller structures. The repeats have a catalytic activity in Peptidyl-glycine alpha-amidating monooxygenase; proteolysis has shown that the Peptidyl-alpha-hydroxyglycine alpha-amidating lyase (PAL) activity is localized to the repeats. Tripartite motif-containing protein 32 interacts with the activation domain of Tat. This interaction is mediated by the NHL repeats. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUU00135.1 | 0.0 | 1 | 998 | 11 | 1005 |
| QBJ17509.1 | 0.0 | 2 | 998 | 11 | 1004 |
| BBK89258.1 | 0.0 | 2 | 998 | 11 | 1004 |
| QUT68914.1 | 0.0 | 2 | 998 | 11 | 1004 |
| QQA31095.1 | 0.0 | 2 | 998 | 11 | 1004 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3EQN_A | 2.62e-10 | 34 | 422 | 53 | 456 | ChainA, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQN_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_A Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D4B0V1 | 1.40e-06 | 369 | 527 | 511 | 666 | Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
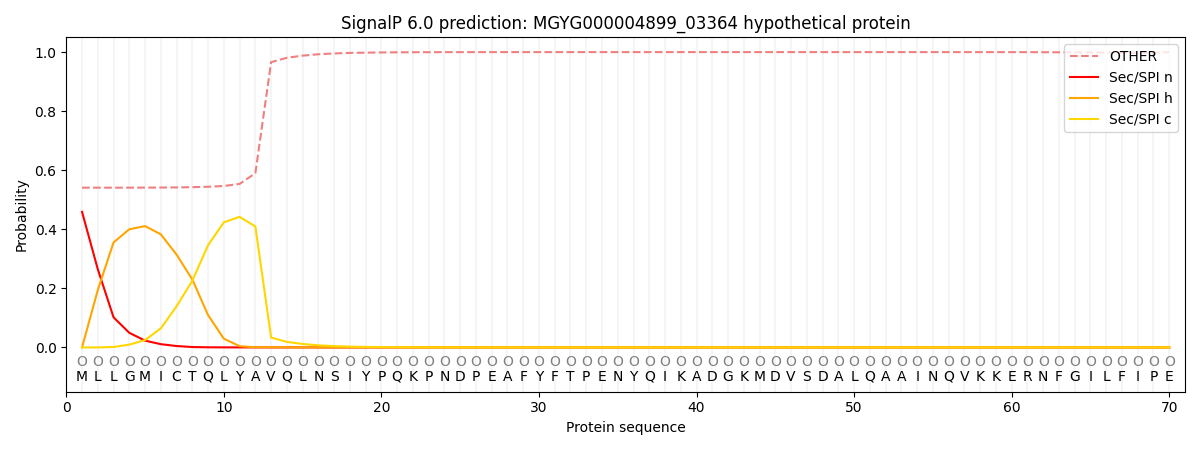
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.556040 | 0.442565 | 0.000345 | 0.000404 | 0.000314 | 0.000339 |
