You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004899_03897
You are here: Home > Sequence: MGYG000004899_03897
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp900555635 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp900555635 | |||||||||||
| CAZyme ID | MGYG000004899_03897 | |||||||||||
| CAZy Family | GH97 | |||||||||||
| CAZyme Description | Retaining alpha-galactosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 26174; End: 28147 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH97 | 11 | 654 | 3.5e-197 | 0.9952456418383518 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam10566 | Glyco_hydro_97 | 3.96e-127 | 282 | 557 | 1 | 278 | Glycoside hydrolase 97. This domain is the catalytic region of the bacterial glycosyl-hydrolase family 97. This central part of the GH97 family protein sequences represents a typical and complete (beta/alpha)8-barrel or catalytic TIM-barrel type domain. The N- and C-terminal parts of the sequences, mainly consisting of beta-strands, form two additional non-catalytic domains. In all known glycosidases with the (beta-alpha)8-barrel fold, the amino acid residues at the active site are located on the C-termini of the beta-strands. |
| pfam14508 | GH97_N | 3.61e-75 | 27 | 277 | 1 | 235 | Glycosyl-hydrolase 97 N-terminal. This N-terminal domain of glycosyl-hydrolase-97 contributes part of the active site pocket. It is also important for contact with the catalytic and C-terminal domains of the whole. |
| pfam14509 | GH97_C | 2.20e-36 | 560 | 655 | 1 | 97 | Glycosyl-hydrolase 97 C-terminal, oligomerization. Glycosyl-hydrolase-97 is made up of three tightly linked and highly conserved globular domains. The C-terminal domain is found to be necessary for oligomerization of the whole molecule in order to create the active-site pocket and the Ca++-binding site. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BBK86458.1 | 0.0 | 2 | 657 | 13 | 668 |
| QPH58760.1 | 0.0 | 1 | 657 | 1 | 657 |
| QUT37383.1 | 0.0 | 2 | 657 | 5 | 660 |
| QUT61529.1 | 0.0 | 2 | 657 | 5 | 660 |
| QQA30035.1 | 0.0 | 2 | 657 | 5 | 660 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3A24_A | 4.58e-97 | 27 | 654 | 6 | 640 | Crystalstructure of BT1871 retaining glycosidase [Bacteroides thetaiotaomicron],3A24_B Crystal structure of BT1871 retaining glycosidase [Bacteroides thetaiotaomicron] |
| 5E1Q_A | 6.91e-96 | 27 | 654 | 20 | 654 | Mutant(D415G) GH97 alpha-galactosidase in complex with Gal-Lac [Bacteroides thetaiotaomicron VPI-5482],5E1Q_B Mutant (D415G) GH97 alpha-galactosidase in complex with Gal-Lac [Bacteroides thetaiotaomicron VPI-5482] |
| 5HQ4_A | 4.52e-83 | 24 | 653 | 2 | 657 | AGlycoside Hydrolase Family 97 enzyme from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8],5HQA_A A Glycoside Hydrolase Family 97 enzyme in complex with Acarbose from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8] |
| 5HQC_A | 4.52e-83 | 24 | 653 | 2 | 657 | AGlycoside Hydrolase Family 97 enzyme R171K variant from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8] |
| 5HQB_A | 1.23e-82 | 24 | 653 | 2 | 657 | AGlycoside Hydrolase Family 97 enzyme (E480Q) in complex with Panose from Pseudoalteromonas sp. strain K8 [Pseudoalteromonas sp. K8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8A6L0 | 3.30e-100 | 1 | 654 | 1 | 661 | Retaining alpha-galactosidase OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=BT_1871 PE=1 SV=1 |
| G8JZS4 | 5.04e-68 | 1 | 654 | 1 | 723 | Glucan 1,4-alpha-glucosidase SusB OS=Bacteroides thetaiotaomicron (strain ATCC 29148 / DSM 2079 / JCM 5827 / CCUG 10774 / NCTC 10582 / VPI-5482 / E50) OX=226186 GN=susB PE=1 SV=1 |
| D7CFN7 | 2.64e-63 | 26 | 649 | 44 | 615 | Probable retaining alpha-galactosidase OS=Streptomyces bingchenggensis (strain BCW-1) OX=749414 GN=SBI_01652 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
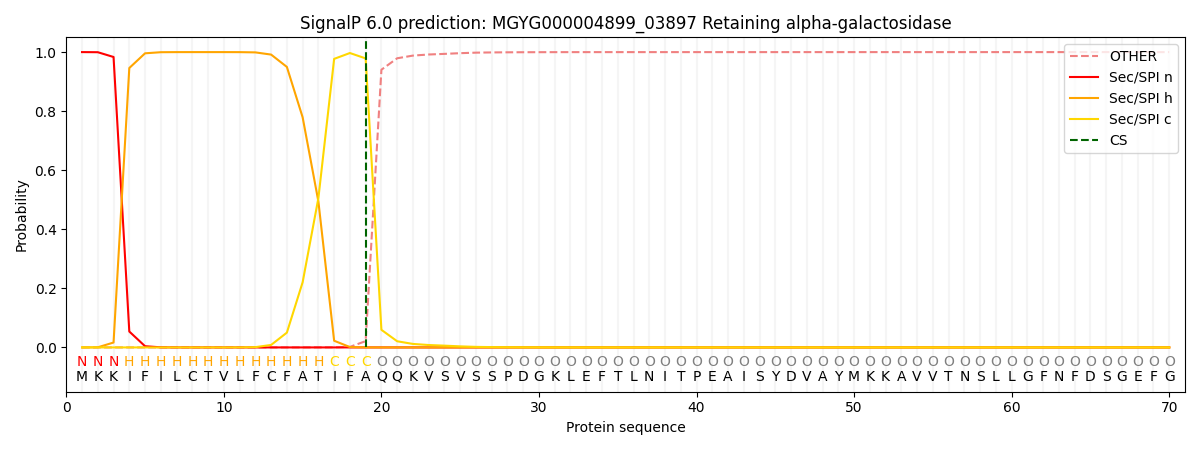
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000299 | 0.998938 | 0.000231 | 0.000175 | 0.000168 | 0.000164 |
