

| Basic Information | |
|---|---|
| Species | Ricinus communis |
| Cazyme ID | 28809.m000040 |
| Family | GH103 |
| Protein Properties | Length: 366 Molecular Weight: 40522.1 Isoelectric Point: 5.0243 |
| Chromosome | Chromosome/Scaffold: 28809 Start: 8201 End: 9301 |
| Description | |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH103 | 2 | 285 | 0 |
| RQEALALHVAPAAVDATLNQVTFLPRLIELDRKQPEFVTSFSAYVNRRINPRVVAKGKQMAEDYQGVLYVVEELYQVPREVLVAFWGLETNFGGFMGDIP LASALATLSYEGRRAEFFKRELLNLMRVVEREQRYDSPVVGSWAGATGHMQFMPSTLLKYGVDADADGKIDIWNSYPDVFSSAANYLSQAGWQPGKPISI QVALPAGFDYSQAQWQLNKPVSEWVKAGVAGVPDSALNLPQAAILLPQGYDGPAYMVFPNFDVIMQWNRSINYALSVSLLSQQL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 366 Download |
| MRQEALALHV APAAVDATLN QVTFLPRLIE LDRKQPEFVT SFSAYVNRRI NPRVVAKGKQ 60 MAEDYQGVLY VVEELYQVPR EVLVAFWGLE TNFGGFMGDI PLASALATLS YEGRRAEFFK 120 RELLNLMRVV EREQRYDSPV VGSWAGATGH MQFMPSTLLK YGVDADADGK IDIWNSYPDV 180 FSSAANYLSQ AGWQPGKPIS IQVALPAGFD YSQAQWQLNK PVSEWVKAGV AGVPDSALNL 240 PQAAILLPQG YDGPAYMVFP NFDVIMQWNR SINYALSVSL LSQQLRQDTY TLMMPPEPPA 300 LSYQQMWSLQ EALNAKGFDC GAPDGFPGAK TQAAIRRYQA SKGLPQDGYA GADIFSRLTA 360 QVQSAQ |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK10760 | PRK10760 | 1.0e-31 | 45 | 275 | 242 | + murein hydrolase B; Provisional | ||
| TIGR02282 | MltB | 3.0e-50 | 13 | 275 | 268 | + lytic murein transglycosylase B. This family consists of lytic murein transglycosylases (murein hydrolases) in the family of MltB, which is a membrane-bound lipoprotein in Escherichia coli. The N-terminal lipoprotein modification motif is conserved in about half the members of this family. The term Slt35 describes a naturally occurring soluble fragment of MltB. Members of this family never contain the putative peptidoglycan binding domain described by pfam01471, which is associated with several classes of bacterial cell wall lytic enzymes [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| COG2951 | MltB | 8.0e-87 | 2 | 275 | 279 | + Membrane-bound lytic murein transglycosylase B [Cell envelope biogenesis, outer membrane] | ||
| TIGR02283 | MltB_2 | 2.0e-98 | 1 | 275 | 281 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| pfam13406 | SLT_2 | 2.0e-119 | 1 | 275 | 276 | + Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002534885.1 | 0 | 1 | 366 | 1 | 366 | Membrane-bound lytic murein transglycosylase B precursor, putative [Ricinus communis] |
| RefSeq | YP_003049020.1 | 0 | 1 | 355 | 39 | 437 | lytic murein transglycosylase [Methylotenera mobilis JLW8] |
| RefSeq | YP_313819.1 | 0 | 1 | 358 | 48 | 407 | lytic murein transglycosylase [Thiobacillus denitrificans ATCC 25259] |
| RefSeq | YP_545322.1 | 0 | 3 | 364 | 38 | 401 | lytic murein transglycosylase [Methylobacillus flagellatus KT] |
| RefSeq | YP_986100.1 | 0 | 3 | 358 | 114 | 471 | lytic murein transglycosylase [Acidovorax sp. JS42] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1qut_A | 1e-28 | 45 | 283 | 78 | 314 | A Chain A, Crystal Structure Of Desr, A Beta-glucosidase From Streptomyces Venezuelae In Complex With D-glucose. |
| PDB | 1qus_A | 1e-28 | 45 | 283 | 78 | 314 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdt_A | 1e-28 | 45 | 283 | 78 | 314 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdr_A | 1e-28 | 45 | 283 | 78 | 314 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1d0m_A | 1e-28 | 45 | 283 | 78 | 314 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO416716 | 274 | 9 | 275 | 0 |
| EX384096 | 284 | 1 | 275 | 0 |
| EX382166 | 230 | 1 | 225 | 0 |
| GO366029 | 230 | 1 | 225 | 0 |
| HO436287 | 200 | 10 | 206 | 1.99965e-42 |
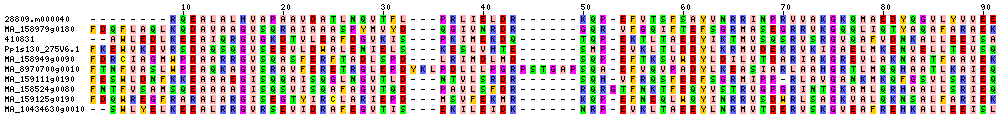
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |