

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_159125g0190 |
| Family | GH103 |
| Protein Properties | Length: 412 Molecular Weight: 45453.6 Isoelectric Point: 10.1309 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH103 | 42 | 333 | 0 |
| FDQWREGFRARALARGISEGTYIRCLARIEPDMSVFEKMRKQPEFNEQLWQYINRRVSDWRLSAGKVALQKNSALFARIEKDFGVERGTLLALWGVETAY GDPLVQQNHMRPVFPSLAALAWNEPRRKAYWETELINALRIVDKHWSTPEEMRGSWAGAMGHTQWMPEVWLNVGFDYDGDGRVSPFGKPDDALGSSARYL VNRGKYRRGEHWGYEVKGATGGSGKKSYAQWAAAGVTRADGQPFPRPNDSAQLWIPVQGGPAFLLGPNFYAVKSYNPSMNYALAIVHLGDRI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 412 Download |
| MTQKDSRNTP ALTRRTLLGS ALGAGALMAS STRALAATPP GFDQWREGFR ARALARGISE 60 GTYIRCLARI EPDMSVFEKM RKQPEFNEQL WQYINRRVSD WRLSAGKVAL QKNSALFARI 120 EKDFGVERGT LLALWGVETA YGDPLVQQNH MRPVFPSLAA LAWNEPRRKA YWETELINAL 180 RIVDKHWSTP EEMRGSWAGA MGHTQWMPEV WLNVGFDYDG DGRVSPFGKP DDALGSSARY 240 LVNRGKYRRG EHWGYEVKGA TGGSGKKSYA QWAAAGVTRA DGQPFPRPND SAQLWIPVQG 300 GPAFLLGPNF YAVKSYNPSM NYALAIVHLG DRILGAGPFI QPFPGSERAL TLAEVQEVQT 360 RLTNAGFDTG GTDGRVGNDT MKAVKDFQTK VGLLPADGYA GLKVLARLRQ GS 420 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK10760 | PRK10760 | 5.0e-22 | 119 | 333 | 221 | + murein hydrolase B; Provisional | ||
| TIGR02282 | MltB | 3.0e-31 | 92 | 333 | 251 | + lytic murein transglycosylase B. This family consists of lytic murein transglycosylases (murein hydrolases) in the family of MltB, which is a membrane-bound lipoprotein in Escherichia coli. The N-terminal lipoprotein modification motif is conserved in about half the members of this family. The term Slt35 describes a naturally occurring soluble fragment of MltB. Members of this family never contain the putative peptidoglycan binding domain described by pfam01471, which is associated with several classes of bacterial cell wall lytic enzymes [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| COG2951 | MltB | 3.0e-88 | 33 | 341 | 317 | + Membrane-bound lytic murein transglycosylase B [Cell envelope biogenesis, outer membrane] | ||
| pfam13406 | SLT_2 | 1.0e-112 | 41 | 331 | 299 | + Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. | ||
| TIGR02283 | MltB_2 | 4.0e-119 | 41 | 335 | 306 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_771020.1 | 0 | 1 | 411 | 1 | 408 | putative transglycolase [Bradyrhizobium japonicum USDA 110] |
| RefSeq | YP_001205585.1 | 0 | 1 | 411 | 1 | 405 | putative membrane-bound lytic transglycosylase precursor protein, putative signal peptide [Bradyrhizobium sp. ORS278] |
| RefSeq | YP_001239979.1 | 0 | 1 | 411 | 1 | 405 | putative membrane-bound lytic transglycosylase precursor protein, putative signal peptide [Bradyrhizobium sp. BTAi1] |
| RefSeq | YP_002289005.1 | 0 | 1 | 411 | 1 | 408 | lytic murein transglycosylase [Oligotropha carboxidovorans OM5] |
| RefSeq | YP_318257.1 | 0 | 1 | 411 | 1 | 406 | lytic murein transglycosylase [Nitrobacter winogradskyi Nb-255] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1ltm_A | 1e-22 | 46 | 333 | 29 | 314 | A Chain A, Accelerated X-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using W |
| PDB | 1qut_A | 1e-22 | 46 | 333 | 31 | 316 | A Chain A, Accelerated X-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using W |
| PDB | 1qus_A | 1e-22 | 46 | 333 | 31 | 316 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdt_A | 1e-22 | 46 | 333 | 31 | 316 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdr_A | 1e-22 | 46 | 333 | 31 | 316 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO416716 | 295 | 51 | 333 | 2e-35 |
| HO436287 | 215 | 56 | 268 | 1e-32 |
| GW436039 | 204 | 214 | 409 | 6e-31 |
| EV111601 | 201 | 210 | 401 | 9e-31 |
| EX384096 | 306 | 45 | 329 | 4e-29 |
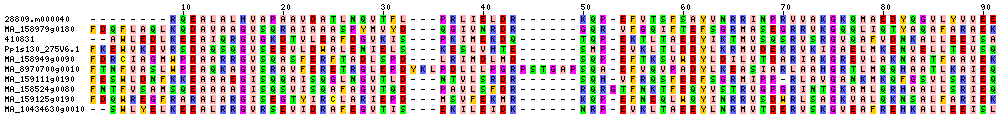
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |