

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_159111g0190 |
| Family | GH103 |
| Protein Properties | Length: 278 Molecular Weight: 30569.6 Isoelectric Point: 8.6654 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH103 | 40 | 244 | 0 |
| FESWLDNFKKEAAAEGISQQAISQGLNGVTLDNTVLSRDRSQHVFRQSFEEFSGRMIPPRLAVGANKMKQFGSVLSRIEQRYGVPGSVLVAIWGLETDYG VNIGKFPTIRSVATLAYDCRRSEMFRAELKDALRIVDQGDLSPTDMRGAWAGEIGQTQFMPSSYIKFAVDFDGNGRRDLLRSAPDVLASTANFLASHGWK RGQPW | |||
| Full Sequence |
|---|
| Protein Sequence Length: 278 Download |
| MFAAGVQMTV GFLLRAAFAA FIVLVPASWA SAATCGTGSF ESWLDNFKKE AAAEGISQQA 60 ISQGLNGVTL DNTVLSRDRS QHVFRQSFEE FSGRMIPPRL AVGANKMKQF GSVLSRIEQR 120 YGVPGSVLVA IWGLETDYGV NIGKFPTIRS VATLAYDCRR SEMFRAELKD ALRIVDQGDL 180 SPTDMRGAWA GEIGQTQFMP SSYIKFAVDF DGNGRRDLLR SAPDVLASTA NFLASHGWKR 240 GQPWDPGSEN FAVIQQWNKS EVYSKTIAYF ANQLDRAP |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR02282 | MltB | 2.0e-50 | 46 | 243 | 207 | + lytic murein transglycosylase B. This family consists of lytic murein transglycosylases (murein hydrolases) in the family of MltB, which is a membrane-bound lipoprotein in Escherichia coli. The N-terminal lipoprotein modification motif is conserved in about half the members of this family. The term Slt35 describes a naturally occurring soluble fragment of MltB. Members of this family never contain the putative peptidoglycan binding domain described by pfam01471, which is associated with several classes of bacterial cell wall lytic enzymes [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| COG2951 | MltB | 4.0e-89 | 12 | 278 | 328 | + Membrane-bound lytic murein transglycosylase B [Cell envelope biogenesis, outer membrane] | ||
| TIGR02283 | MltB_2 | 5.0e-104 | 39 | 276 | 300 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| pfam13406 | SLT_2 | 7.0e-106 | 39 | 244 | 207 | + Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_771364.1 | 0 | 31 | 278 | 27 | 274 | hypothetical protein blr4724 [Bradyrhizobium japonicum USDA 110] |
| RefSeq | YP_001206000.1 | 0 | 8 | 278 | 1 | 271 | putative transglycolase [Bradyrhizobium sp. ORS278] |
| RefSeq | YP_001240341.1 | 0 | 13 | 278 | 6 | 271 | putative transglycolase [Bradyrhizobium sp. BTAi1] |
| RefSeq | YP_001834412.1 | 0 | 23 | 275 | 26 | 280 | lytic murein transglycosylase [Beijerinckia indica subsp. indica ATCC 9039] |
| RefSeq | ZP_05376432.1 | 0 | 14 | 274 | 13 | 277 | lytic murein transglycosylase [Hyphomicrobium denitrificans ATCC 51888] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 4anr_A | 3e-27 | 99 | 243 | 82 | 227 | A Chain A, Crystal Structure Of Soluble Lytic Transglycosylase Sltb1 From Pseudomonas Aeruginosa |
| PDB | 1qut_A | 3e-24 | 95 | 241 | 83 | 229 | A Chain A, Crystal Structure Of Soluble Lytic Transglycosylase Sltb1 From Pseudomonas Aeruginosa |
| PDB | 1qus_A | 3e-24 | 95 | 241 | 83 | 229 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdt_A | 3e-24 | 95 | 241 | 83 | 229 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdr_A | 3e-24 | 95 | 241 | 83 | 229 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| Transmembrane Domains | ||||
|---|---|---|---|---|
 | ||||
| Start | End | |||
| 12 | 34 | |||
| Signal Peptide | ||||
|---|---|---|---|---|
 | ||||
| Cleavage Site | ||||
| 0 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO436287 | 194 | 55 | 244 | 0 |
| EX384096 | 212 | 39 | 243 | 0 |
| EX382166 | 212 | 39 | 243 | 0 |
| GO366029 | 212 | 39 | 243 | 0 |
| HO416716 | 194 | 56 | 244 | 5e-39 |
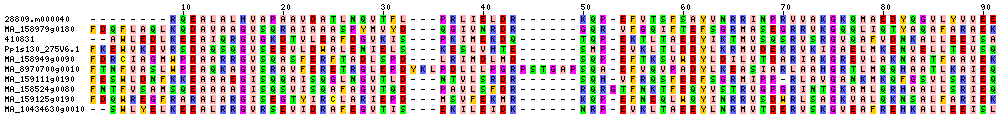
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |