

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_158524g0080 |
| Family | GH103 |
| Protein Properties | Length: 269 Molecular Weight: 29257.4 Isoelectric Point: 9.8085 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH103 | 29 | 235 | 0 |
| FNTFVSAMSQEAAAAGISQSVISQAFAGVTQDPAVLSFDRRQRGTFNKTFEQYVSTRVGPGRINTGKAMLQRHAALLSRIEQRFGVPPEIVVAIWGLESD FGKGDIGKMPVIRTLTTLAHDCRRTELFQGELLAALKIVQRGDLPLRDLIGAYAGELGQTQFLPSSYIKYGVDFDGNGHVDLRHSVPDVLASTANLLKTA GFRGGAD | |||
| Full Sequence |
|---|
| Protein Sequence Length: 269 Download |
| MMFRLRLAVI AAALSLAAPA NAARCGGDFN TFVSAMSQEA AAAGISQSVI SQAFAGVTQD 60 PAVLSFDRRQ RGTFNKTFEQ YVSTRVGPGR INTGKAMLQR HAALLSRIEQ RFGVPPEIVV 120 AIWGLESDFG KGDIGKMPVI RTLTTLAHDC RRTELFQGEL LAALKIVQRG DLPLRDLIGA 180 YAGELGQTQF LPSSYIKYGV DFDGNGHVDL RHSVPDVLAS TANLLKTAGF RGGADYHEGS 240 ANFEAMREWN RATIYRKTIA YFADRLAGR |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR02283 | MltB_2 | 3.0e-7 | 215 | 268 | 54 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| COG2951 | MltB | 9.0e-83 | 25 | 263 | 243 | + Membrane-bound lytic murein transglycosylase B [Cell envelope biogenesis, outer membrane] | ||
| pfam13406 | SLT_2 | 3.0e-100 | 28 | 236 | 209 | + Transglycosylase SLT domain. This family is related to the SLT domain pfam01464. | ||
| TIGR02283 | MltB_2 | 8.0e-102 | 28 | 233 | 207 | + lytic murein transglycosylase. Members of this family are closely related to the MltB family lytic murein transglycosylases described by TIGR02282 and are likewise all proteobacterial, although that family and this one form clearly distinct clades. Several species have one member of each family. Many members of this family (unlike the MltB family) contain an additional C-terminal domain, a putative peptidoglycan binding domain (pfam01471), not included in region described by this model. Many sequences appear to contain N-terminal lipoprotein attachment sites, as does E. coli MltB in TIGR02282 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan]. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_773984.1 | 0 | 2 | 269 | 1 | 268 | hypothetical protein bll7344 [Bradyrhizobium japonicum USDA 110] |
| RefSeq | NP_946654.1 | 0 | 2 | 266 | 1 | 269 | lytic murein transglycosylase [Rhodopseudomonas palustris CGA009] |
| RefSeq | YP_571085.1 | 0 | 24 | 269 | 27 | 272 | lytic murein transglycosylase [Rhodopseudomonas palustris BisB5] |
| RefSeq | YP_779977.1 | 0 | 24 | 267 | 23 | 266 | lytic murein transglycosylase [Rhodopseudomonas palustris BisA53] |
| RefSeq | ZP_06358932.1 | 0 | 2 | 266 | 1 | 269 | lytic murein transglycosylase [Rhodopseudomonas palustris DX-1] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 4anr_A | 6e-25 | 32 | 233 | 28 | 225 | A Chain A, Crystal Structure Of Soluble Lytic Transglycosylase Sltb1 From Pseudomonas Aeruginosa |
| PDB | 1ltm_A | 3e-24 | 75 | 233 | 70 | 227 | A Chain A, Accelerated X-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using W |
| PDB | 1qut_A | 3e-24 | 75 | 233 | 72 | 229 | A Chain A, Accelerated X-ray Structure Elucidation Of A 36 Kda Muramidase/transglycosylase Using W |
| PDB | 1qus_A | 3e-24 | 75 | 233 | 72 | 229 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| PDB | 1qdt_A | 3e-24 | 75 | 233 | 72 | 229 | A Chain A, 1.7 A Resolution Structure Of The Soluble Lytic Transglycosylase Slt35 From Escherichia Coli |
| Signal Peptide | ||||
|---|---|---|---|---|
 | ||||
| Cleavage Site | ||||
| 0 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO436287 | 217 | 44 | 257 | 1.99993e-41 |
| EX382166 | 209 | 30 | 233 | 4e-36 |
| EX384096 | 209 | 30 | 233 | 4e-36 |
| GO366029 | 209 | 30 | 233 | 4e-36 |
| HO416716 | 196 | 45 | 236 | 1e-32 |
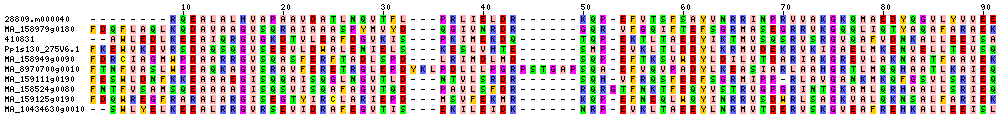
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |